Krüppel-like factor 10 modulates stem cell phenotypes of pancreatic adenocarcinoma by transcriptionally regulating notch receptors
- PMID: 37308977
- PMCID: PMC10258947
- DOI: 10.1186/s12929-023-00937-z
Krüppel-like factor 10 modulates stem cell phenotypes of pancreatic adenocarcinoma by transcriptionally regulating notch receptors
Abstract
Background: Pancreatic adenocarcinoma (PDAC) is well known for its rapid distant metastasis and local destructive behavior. Loss of Krüppel-like factor 10 (KLF10) contributes to distant migration of PDAC. The role of KLF10 in modulating tumorigenesis and stem cell phenotypes of PDAC is unclear.
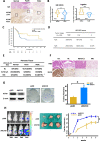
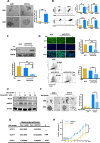
Methods: Additional depletion of KLF10 in KC (LSL: KrasG12D; Pdx1-Cre) mice, a spontaneous murine PDAC model, was established to evaluate tumorigenesis. Tumor specimens of PDAC patients were immune-stained of KLF10 to correlate with local recurrence after curative resection. Conditional overexpressing KLF10 in MiaPaCa and stably depleting KLF10 in Panc-1 (Panc-1-pLKO-shKLF10) cells were established for evaluating sphere formation, stem cell markers expression and tumor growth. The signal pathways modulated by KLF10 for PDAC stem cell phenotypes were disclosed by microarray analysis and validated by western blot, qRT-PCR, luciferase reporter assay. Candidate targets to reverse PDAC tumor growth were demonstrated in murine model.
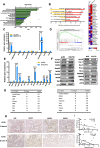
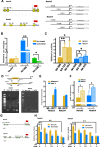
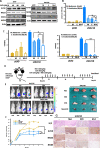
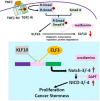
Results: KLF10, deficient in two-thirds of 105 patients with resected pancreatic PDAC, was associated with rapid local recurrence and large tumor size. Additional KLF10 depletion in KC mice accelerated progression from pancreatic intraepithelial neoplasia to PDAC. Increased sphere formation, expression of stem cell markers, and tumor growth were observed in Panc-1-pLKO-shKLF10 compared with vector control. Genetically or pharmacologically overexpression of KLF10 reversed the stem cell phenotypes induced by KLF10 depletion. Ingenuity pathway analysis and gene set enrichment analysis showed that Notch signaling molecules, including Notch receptors 3 and 4, were over-expressed in Panc-1-pLKO-shKLF10. KLF10 transcriptionally suppressed Notch-3 and -4 by competing with E74-like ETS transcription factor 3, a positive regulator, for promoter binding. Downregulation of Notch signaling, either genetically or pharmacologically, ameliorated the stem cell phenotypes of Panc-1-pLKO-shKLF10. The combination of metformin, which upregulated KLF10 expression via phosphorylating AMPK, and evodiamine, a non-toxic Notch-3 methylation stimulator, delayed tumor growth of PDAC with KLF10 deficiency in mice without prominent toxicity.
Conclusions: These results demonstrated a novel signaling pathway by which KLF10 modulates stem cell phenotypes in PDAC through transcriptionally regulating Notch signaling pathway. The elevation of KLF10 and suppression of Notch signaling may jointly reduce PDAC tumorigenesis and malignant progression.
Keywords: ELF3; Krüppel-like factor 10; Notch signaling; Notch-3/4; Pancreatic adenocarcinoma.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Krüppel-like Factor 10 as a Prognostic and Predictive Biomarker of Radiotherapy in Pancreatic Adenocarcinoma.Cancers (Basel). 2023 Oct 30;15(21):5212. doi: 10.3390/cancers15215212. Cancers (Basel). 2023. PMID: 37958386 Free PMC article. Review.
-
Upregulating sirtuin 6 ameliorates glycolysis, EMT and distant metastasis of pancreatic adenocarcinoma with krüppel-like factor 10 deficiency.Exp Mol Med. 2021 Oct;53(10):1623-1635. doi: 10.1038/s12276-021-00687-8. Epub 2021 Oct 26. Exp Mol Med. 2021. PMID: 34702956 Free PMC article.
-
Lunatic Fringe is a potent tumor suppressor in Kras-initiated pancreatic cancer.Oncogene. 2016 May 12;35(19):2485-95. doi: 10.1038/onc.2015.306. Epub 2015 Aug 17. Oncogene. 2016. PMID: 26279302
-
Krüpple-like factor 10 regulates radio-sensitivity of pancreatic cancer via UV radiation resistance-associated gene.Radiother Oncol. 2017 Mar;122(3):476-484. doi: 10.1016/j.radonc.2017.01.001. Epub 2017 Jan 16. Radiother Oncol. 2017. PMID: 28104298
-
Notch signaling pathway in pancreatic tumorigenesis.Adv Cancer Res. 2023;159:1-36. doi: 10.1016/bs.acr.2023.02.001. Epub 2023 Feb 28. Adv Cancer Res. 2023. PMID: 37268393 Review.
Cited by
-
FXYD6 is transcriptionally activated by KLF10 to suppress the aggressiveness of gastric cancer cells.Cytotechnology. 2025 Apr;77(2):48. doi: 10.1007/s10616-025-00710-0. Epub 2025 Jan 25. Cytotechnology. 2025. PMID: 39867832
-
The Interplay between Medical Plants and Gut Microbiota in Cancer.Nutrients. 2023 Jul 26;15(15):3327. doi: 10.3390/nu15153327. Nutrients. 2023. PMID: 37571264 Free PMC article. Review.
-
Krüppel-like Factor 10 as a Prognostic and Predictive Biomarker of Radiotherapy in Pancreatic Adenocarcinoma.Cancers (Basel). 2023 Oct 30;15(21):5212. doi: 10.3390/cancers15215212. Cancers (Basel). 2023. PMID: 37958386 Free PMC article. Review.
-
Notch signaling pathway in cancer: from mechanistic insights to targeted therapies.Signal Transduct Target Ther. 2024 May 27;9(1):128. doi: 10.1038/s41392-024-01828-x. Signal Transduct Target Ther. 2024. PMID: 38797752 Free PMC article. Review.
References
-
- Ercan G, Karlitepe A, Ozpolat B. Pancreatic cancer stem cells and therapeutic approaches. Anticancer Res. 2017;37(6):2761–2775. - PubMed
-
- Weng CC, Hawse JR, Subramaniam M, Chang VHS, Yu WCY, Hung WC, Chen LT, Cheng KH. KLF10 loss in the pancreas provokes activation of SDF-1 and induces distant metastases of pancreatic ductal adenocarcinoma in the Kras(G12D) p53(flox/flox) model. Oncogene. 2017;36(39):5532–5543. doi: 10.1038/onc.2017.155. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

