SIRT1 regulates DNA damage signaling through the PP4 phosphatase complex
- PMID: 37309898
- PMCID: PMC10359614
- DOI: 10.1093/nar/gkad504
SIRT1 regulates DNA damage signaling through the PP4 phosphatase complex
Abstract
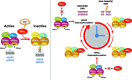
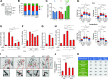
The Sirtuin family of NAD+-dependent enzymes plays an important role in maintaining genome stability upon stress. Several mammalian Sirtuins have been linked directly or indirectly to the regulation of DNA damage during replication through Homologous recombination (HR). The role of one of them, SIRT1, is intriguing as it seems to have a general regulatory role in the DNA damage response (DDR) that has not yet been addressed. SIRT1-deficient cells show impaired DDR reflected in a decrease in repair capacity, increased genome instability and decreased levels of γH2AX. Here we unveil a close functional antagonism between SIRT1 and the PP4 phosphatase multiprotein complex in the regulation of the DDR. Upon DNA damage, SIRT1 interacts specifically with the catalytical subunit PP4c and promotes its inhibition by deacetylating the WH1 domain of the regulatory subunits PP4R3α/β. This in turn regulates γH2AX and RPA2 phosphorylation, two key events in the signaling of DNA damage and repair by HR. We propose a mechanism whereby during stress, SIRT1 signaling ensures a global control of DNA damage signaling through PP4.
© The Author(s) 2023. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures






References
-
- Guarente L. Franklin H. Epstein Lecture: sirtuins, aging, and medicine. N. Engl. J. Med. 2011; 364:2235–2244. - PubMed
-
- Bosch-Presegue L., Vaquero A.. Sirtuin-dependent epigenetic regulation in the maintenance of genome integrity. FEBS J. 2015; 282:1745–1767. - PubMed
-
- Sedelnikova O.A., Rogakou E.P., Panyutin I.G., Bonner W.M.. Quantitative detection of (125)IdU-induced DNA double-strand breaks with gamma-H2AX antibody. Radiat. Res. 2002; 158:486–492. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

