A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
- PMID: 37316492
- PMCID: PMC10267216
- DOI: 10.1038/s41467-023-38902-x
A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
Abstract
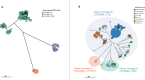
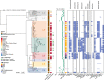
Antimicrobial resistant Salmonella enterica serovar Concord (S. Concord) is known to cause severe gastrointestinal and bloodstream infections in patients from Ethiopia and Ethiopian adoptees, and occasional records exist of S. Concord linked to other countries. The evolution and geographical distribution of S. Concord remained unclear. Here, we provide a genomic overview of the population structure and antimicrobial resistance (AMR) of S. Concord by analysing genomes from 284 historical and contemporary isolates obtained between 1944 and 2022 across the globe. We demonstrate that S. Concord is a polyphyletic serovar distributed among three Salmonella super-lineages. Super-lineage A is composed of eight S. Concord lineages, of which four are associated with multiple countries and low levels of AMR. Other lineages are restricted to Ethiopia and horizontally acquired resistance to most antimicrobials used for treating invasive Salmonella infections in low- and middle-income countries. By reconstructing complete genomes for 10 representative strains, we demonstrate the presence of AMR markers integrated in structurally diverse IncHI2 and IncA/C2 plasmids, and/or the chromosome. Molecular surveillance of pathogens such as S. Concord supports the understanding of AMR and the multi-sector response to the global AMR threat. This study provides a comprehensive baseline data set essential for future molecular surveillance.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
References
-
- Grimont PAD, W. F. X. Antigenic formulae of the Salmonella serovars, 9th ed. (World health organization collaborating center for reference and research on Salmonella, Insitut Pasteur, 2007).
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
- BBS/E/F/000PR10349/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/R012504/1 /BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- MC_PC_16093/MRC_/Medical Research Council/United Kingdom
- BB/CCG1720/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- WT_/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

