Osteoarthritic chondrocytes undergo a glycolysis-related metabolic switch upon exposure to IL-1b or TNF
- PMID: 37316888
- PMCID: PMC10265918
- DOI: 10.1186/s12964-023-01150-z
Osteoarthritic chondrocytes undergo a glycolysis-related metabolic switch upon exposure to IL-1b or TNF
Abstract
Background: Osteoarthritis is an age-related disease that currently faces a lack of symptomatic treatment. Inflammation, which is mainly sustained by pro-inflammatory cytokines such as IL-1b, TNF, and IL-6, plays an important role in osteoarthritis progression. In this context, pro-inflammatory cytokines are widely used to mimic the inflammatory component of osteoarthritis in vitro. However, the therapeutic failures of clinical trials evaluating anti-cytokines drugs highlight the lack of overall understanding of the effects of these cytokines on chondrocytes.
Methods: Here, we generated a comprehensive transcriptomic and proteomic dataset of osteoarthritic chondrocytes treated with these cytokines to describe their pro-inflammatory signature and compare it to the transcriptome of non-osteoarthritic chondrocytes. Then, the dysregulations highlighted at the molecular level were functionally confirmed by real-time cellular metabolic assays.
Results: We identified dysregulation of metabolic-related genes in osteoarthritic chondrocytes but not in non-osteoarthritic chondrocytes. A metabolic shift, toward increased glycolysis at the expense of mitochondrial respiration, was specifically confirmed in osteoarthritic chondrocytes treated with IL-1b or TNF.
Conclusion: These data show a strong and specific association between inflammation and metabolism in osteoarthritic chondrocytes, which was not found in non-osteoarthritic chondrocytes. This indicates that the link between inflammation and metabolic dysregulation may be exacerbated during chondrocyte damage in osteoarthritis. Video Abstract.
Keywords: Chondrocytes; Inflammation; Metabolism; Osteoarthritis.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

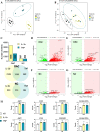




References
-
- Hunter DJ, Bierma-Zeinstra S. Osteoarthritis Lancet. 2019;393:1745–1759. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

