Global translational control by the transcriptional repressor TrcR in the filamentous cyanobacterium Anabaena sp. PCC 7120
- PMID: 37322092
- PMCID: PMC10272220
- DOI: 10.1038/s42003-023-05012-9
Global translational control by the transcriptional repressor TrcR in the filamentous cyanobacterium Anabaena sp. PCC 7120
Abstract
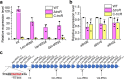
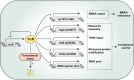
Transcriptional and translational regulations are important mechanisms for cell adaptation to environmental conditions. In addition to house-keeping tRNAs, the genome of the filamentous cyanobacterium Anabaena sp. strain PCC 7120 (Anabaena) has a long tRNA operon (trn operon) consisting of 26 genes present on a megaplasmid. The trn operon is repressed under standard culture conditions, but is activated under translational stress in the presence of antibiotics targeting translation. Using the toxic amino acid analog β-N-methylamino-L-alanine (BMAA) as a tool, we isolated and characterized several BMAA-resistance mutants from Anabaena, and identified one gene of unknown function, all0854, named as trcR, encoding a transcription factor belonging to the ribbon-helix-helix (RHH) family. We provide evidence that TrcR represses the expression of the trn operon and is thus the missing link between the trn operon and translational stress response. TrcR represses the expression of several other genes involved in translational control, and is required for maintaining translational fidelity. TrcR, as well as its binding sites, are highly conserved in cyanobacteria, and its functions represent an important mechanism for the coupling of the transcriptional and translational regulations in cyanobacteria.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







Similar articles
-
A tRNA t6A modification system contributes to the sensitivity towards the toxin β-N-methylamino-L-alanine (BMAA) in the cyanobacterium Anabaena sp. PCC 7120.Aquat Toxicol. 2022 Apr;245:106121. doi: 10.1016/j.aquatox.2022.106121. Epub 2022 Feb 13. Aquat Toxicol. 2022. PMID: 35180454
-
Heterocyst Septa Contain Large Nanopores That Are Influenced by the Fra Proteins in the Filamentous Cyanobacterium Anabaena sp. Strain PCC 7120.J Bacteriol. 2021 Jun 8;203(13):e0008121. doi: 10.1128/JB.00081-21. Epub 2021 Jun 8. J Bacteriol. 2021. PMID: 33846119 Free PMC article.
-
Induction of the Nitrate Assimilation nirA Operon and Protein-Protein Interactions in the Maturation of Nitrate and Nitrite Reductases in the Cyanobacterium Anabaena sp. Strain PCC 7120.J Bacteriol. 2015 Jul;197(14):2442-52. doi: 10.1128/JB.00198-15. Epub 2015 May 11. J Bacteriol. 2015. PMID: 25962912 Free PMC article.
-
ATP-binding cassette transporters of the multicellular cyanobacterium Anabaena sp. PCC 7120: a wide variety for a complex lifestyle.FEMS Microbiol Lett. 2018 Feb 1;365(4). doi: 10.1093/femsle/fny012. FEMS Microbiol Lett. 2018. PMID: 29360977 Review.
-
Genetic tools for cyanobacteria.Appl Microbiol Biotechnol. 2002 Feb;58(2):123-37. doi: 10.1007/s00253-001-0864-9. Appl Microbiol Biotechnol. 2002. PMID: 11876404 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

