Cellulose metabolism in halo(natrono)archaea: a comparative genomics study
- PMID: 37323904
- PMCID: PMC10267330
- DOI: 10.3389/fmicb.2023.1112247
Cellulose metabolism in halo(natrono)archaea: a comparative genomics study
Abstract
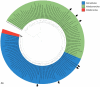
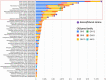
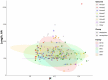
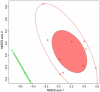
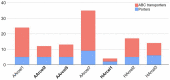
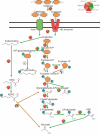
Extremely halophilic archaea are one of the principal microbial community components in hypersaline environments. The majority of cultivated haloarchaea are aerobic heterotrophs using peptides or simple sugars as carbon and energy sources. At the same time, a number of novel metabolic capacities of these extremophiles were discovered recently among which is a capability of growing on insoluble polysaccharides such as cellulose and chitin. Still, polysaccharidolytic strains are in minority among cultivated haloarchaea and their capacities of hydrolyzing recalcitrant polysaccharides are hardly investigated. This includes the mechanisms and enzymes involved in cellulose degradation, which are well studied for bacterial species, while almost unexplored in archaea and haloarchaea in particular. To fill this gap, a comparative genomic analysis of 155 cultivated representatives of halo(natrono)archaea, including seven cellulotrophic strains belonging to the genera Natronobiforma, Natronolimnobius, Natrarchaeobius, Halosimplex, Halomicrobium and Halococcoides was performed. The analysis revealed a number of cellulases, encoded in the genomes of cellulotrophic strains but also in several haloarchaea, for which the capacity to grow on cellulose was not shown. Surprisingly, the cellulases genes, especially of GH5, GH9 and GH12 families, were significantly overrepresented in the cellulotrophic haloarchaea genomes in comparison with other cellulotrophic archaea and even cellulotrophic bacteria. Besides cellulases, the genes for GH10 and GH51 families were also abundant in the genomes of cellulotrophic haloarchaea. These results allowed to propose the genomic patterns, determining the capability of haloarchaea to grow on cellulose. The patterns helped to predict cellulotrophic capacity for several halo(natrono)archaea, and for three of them it was experimentally confirmed. Further genomic search revealed that glucose and cellooligosaccharides import occurred by means of porters and ABC (ATP-binding cassette) transporters. Intracellular glucose oxidation occurred through glycolysis or the semi-phosphorylative Entner-Dudoroff pathway which occurrence was strain-specific. Comparative analysis of CAZymes toolbox and available cultivation-based information allowed proposing two possible strategies used by haloarchaea capable of growing on cellulose: so-called specialists are more effective in degradation of cellulose while generalists are more flexible in nutrient spectra. Besides CAZymes profiles the groups differed in genome sizes, as well as in variability of mechanisms of import and central metabolism of sugars.
Keywords: CAZymes; cellulose; cellulotrophic; genomics; haloarchaea; polysaccharides degradation.
Copyright © 2023 Elcheninov, Ugolkov, Elizarov, Klyukina, Kublanov and Sorokin.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Isolation and characterization of cellulose-mineralizing haloalkaliphilic bacteria from Siberian soda lakes.Front Microbiol. 2024 Dec 23;15:1523074. doi: 10.3389/fmicb.2024.1523074. eCollection 2024. Front Microbiol. 2024. PMID: 39764452 Free PMC article.
-
Halo(natrono)archaea isolated from hypersaline lakes utilize cellulose and chitin as growth substrates.Front Microbiol. 2015 Sep 15;6:942. doi: 10.3389/fmicb.2015.00942. eCollection 2015. Front Microbiol. 2015. PMID: 26441877 Free PMC article.
-
Selective enrichment on a wide polysaccharide spectrum allowed isolation of novel metabolic and taxonomic groups of haloarchaea from hypersaline lakes.Front Microbiol. 2022 Nov 23;13:1059347. doi: 10.3389/fmicb.2022.1059347. eCollection 2022. Front Microbiol. 2022. PMID: 36504804 Free PMC article.
-
Cellulolytic Aerobic Bacteria Isolated from Agricultural and Forest Soils: An Overview.Biology (Basel). 2024 Feb 5;13(2):102. doi: 10.3390/biology13020102. Biology (Basel). 2024. PMID: 38392320 Free PMC article. Review.
-
Polyhydroxyalkanoate Biosynthesis at the Edge of Water Activitiy-Haloarchaea as Biopolyester Factories.Bioengineering (Basel). 2019 Apr 16;6(2):34. doi: 10.3390/bioengineering6020034. Bioengineering (Basel). 2019. PMID: 30995811 Free PMC article. Review.
Cited by
-
The Isolation and Characterization of a Novel Psychrotolerant Cellulolytic Bacterium, Microbacterium sp. QXD-8T.Microorganisms. 2024 Jan 31;12(2):303. doi: 10.3390/microorganisms12020303. Microorganisms. 2024. PMID: 38399707 Free PMC article.
-
Natronoglomus mannanivorans gen. nov., sp. nov., beta-1,4-mannan utilizing natronoarchaea from hypersaline soda lakes.Front Microbiol. 2024 Mar 12;15:1364606. doi: 10.3389/fmicb.2024.1364606. eCollection 2024. Front Microbiol. 2024. PMID: 38533326 Free PMC article.
-
Natrarchaeobius versutus sp. nov. and Natrarchaeobius oligotrophus sp. nov., chitinotrophic natronoarchaea from hypersaline soda lakes, and functional genome analysis of the Natrarchaeobius species.Front Microbiol. 2025 Jul 30;16:1640521. doi: 10.3389/fmicb.2025.1640521. eCollection 2025. Front Microbiol. 2025. PMID: 40809049 Free PMC article.
-
Isolation and characterization of cellulose-mineralizing haloalkaliphilic bacteria from Siberian soda lakes.Front Microbiol. 2024 Dec 23;15:1523074. doi: 10.3389/fmicb.2024.1523074. eCollection 2024. Front Microbiol. 2024. PMID: 39764452 Free PMC article.
References
-
- Altekar W., Rangaswamy V. (1991). Ketohexokinase (ATP: D-fructose 1-phosphotransferase) initiates fructose breakdown via the modified EMP pathway in halophilic archaebacteria. FEMS Microbiol. Lett. 83, 241–246. doi: 10.1111/j.1574-6968.1991.tb04471.x - DOI
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

