Functional diversity of nanohaloarchaea within xylan-degrading consortia
- PMID: 37323909
- PMCID: PMC10266531
- DOI: 10.3389/fmicb.2023.1182464
Functional diversity of nanohaloarchaea within xylan-degrading consortia
Abstract
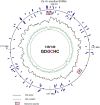
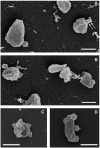
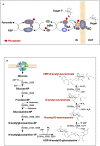
Extremely halophilic representatives of the phylum Candidatus Nanohaloarchaeota (members of the DPANN superphyla) are obligately associated with extremely halophilic archaea of the phylum Halobacteriota (according to the GTDB taxonomy). Using culture-independent molecular techniques, their presence in various hypersaline ecosystems around the world has been confirmed over the past decade. However, the vast majority of nanohaloarchaea remain uncultivated, and thus their metabolic capabilities and ecophysiology are currently poorly understood. Using the (meta)genomic, transcriptomic, and DNA methylome platforms, the metabolism and functional prediction of the ecophysiology of two novel extremely halophilic symbiotic nanohaloarchaea (Ca. Nanohalococcus occultus and Ca. Nanohalovita haloferacivicina) stably cultivated in the laboratory as members of a xylose-degrading binary culture with a haloarchaeal host, Haloferax lucentense, was determined. Like all known DPANN superphylum nanoorganisms, these new sugar-fermenting nanohaloarchaea lack many fundamental biosynthetic repertoires, making them exclusively dependent on their respective host for survival. In addition, given the cultivability of the new nanohaloarchaea, we managed to discover many unique features in these new organisms that have never been observed in nano-sized archaea both within the phylum Ca. Nanohaloarchaeota and the entire superphylum DPANN. This includes the analysis of the expression of organism-specific non-coding regulatory (nc)RNAs (with an elucidation of their 2D-secondary structures) as well as profiling of DNA methylation. While some ncRNA molecules have been predicted with high confidence as RNAs of an archaeal signal recognition particle involved in delaying protein translation, others resemble the structure of ribosome-associated ncRNAs, although none belong to any known family. Moreover, the new nanohaloarchaea have very complex cellular defense mechanisms. In addition to the defense mechanism provided by the type II restriction-modification system, consisting of Dcm-like DNA methyltransferase and Mrr restriction endonuclease, Ca. Nanohalococcus encodes an active type I-D CRISPR/Cas system, containing 77 spacers divided into two loci. Despite their diminutive genomes and as part of their host interaction mechanism, the genomes of new nanohaloarchaea do encode giant surface proteins, and one of them (9,409 amino acids long) is the largest protein of any sequenced nanohaloarchaea and the largest protein ever discovered in cultivated archaea.
Keywords: CRISPR; ecology of nanohaloarchaea; methylomics; nanohaloarchaeal-haloarchaeal symbioses; ncRNA.
Copyright © 2023 Reva, Messina, La Cono, Crisafi, Smedile, La Spada, Marturano, Selivanova, Rohde, Krupovic and Yakimov.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
LinkOut - more resources
Full Text Sources
Miscellaneous

