What remains from living cells in bacterial lysate-based cell-free systems
- PMID: 37333859
- PMCID: PMC10275740
- DOI: 10.1016/j.csbj.2023.05.025
What remains from living cells in bacterial lysate-based cell-free systems
Abstract
Because they mimic cells while offering an accessible and controllable environment, lysate-based cell-free systems (CFS) have emerged as valuable biotechnology tools for synthetic biology. Historically used to uncover fundamental mechanisms of life, CFS are nowadays used for a multitude of purposes, including protein production and prototyping of synthetic circuits. Despite the conservation of fundamental functions in CFS like transcription and translation, RNAs and certain membrane-embedded or membrane-bound proteins of the host cell are lost when preparing the lysate. As a result, CFS largely lack some essential properties of living cells, such as the ability to adapt to changing conditions, to maintain homeostasis and spatial organization. Regardless of the application, shedding light on the black-box of the bacterial lysate is necessary to fully exploit the potential of CFS. Most measurements of the activity of synthetic circuits in CFS and in vivo show significant correlations because these only require processes that are preserved in CFS, like transcription and translation. However, prototyping circuits of higher complexity that require functions that are lost in CFS (cell adaptation, homeostasis, spatial organization) will not show such a good correlation with in vivo conditions. Both for prototyping circuits of higher complexity and for building artificial cells, the cell-free community has developed devices to reconstruct cellular functions. This mini-review compares bacterial CFS to living cells, focusing on functional and cellular process differences and the latest developments in restoring lost functions through complementation of the lysate or device engineering.
Keywords: Adaptation; Cell-free; E. coli; Homeostasis; Microfluidics; Prototyping; Spatial organization.
© 2023 The Authors. Published by Elsevier B.V. on behalf of Research Network of Computational and Structural Biotechnology.
Conflict of interest statement
The authors declare that they have no affiliations with or involvement in any organization or entity with any financial interest in the subject matter or materials discussed in this manuscript.
Figures

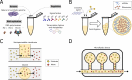
References
-
- Garenne D., Haines M.C., Romantseva E.F., Freemont P., Strychalski E.A., Noireaux V. Cell-free gene expression. Nat Rev Methods Prim. 2021;1:1–18. doi: 10.1038/s43586-021-00046-x. - DOI
Publication types
LinkOut - more resources
Full Text Sources
Research Materials
