Glyoxylate Shunt and Pyruvate-to-Acetoin Shift Are Specific Stress Responses Induced by Colistin and Ceragenin CSA-13 in Enterobacter hormaechei ST89
- PMID: 37338344
- PMCID: PMC10434160
- DOI: 10.1128/spectrum.01215-23
Glyoxylate Shunt and Pyruvate-to-Acetoin Shift Are Specific Stress Responses Induced by Colistin and Ceragenin CSA-13 in Enterobacter hormaechei ST89
Abstract
Ceragenins, including CSA-13, are cationic antimicrobials that target the bacterial cell envelope differently than colistin. However, the molecular basis of their action is not fully understood. Here, we examined the genomic and transcriptome responses by Enterobacter hormaechei after prolonged exposure to either CSA-13 or colistin. Resistance of the E. hormaechei 4236 strain (sequence type 89 [ST89]) to colistin and CSA-13 was induced in vitro during serial passages with sublethal doses of tested agents. The genomic and metabolic profiles of the tested isolates were characterized using a combination of whole-genome sequencing (WGS) and transcriptome sequencing (RNA-seq), followed by metabolic mapping of differentially expressed genes using Pathway Tools software. The exposure of E. hormaechei to colistin resulted in the deletion of the mgrB gene, whereas CSA-13 disrupted the genes encoding an outer membrane protein C and transcriptional regulator SmvR. Both compounds upregulated several colistin-resistant genes, such as the arnABCDEF operon and pagE, including genes coding for DedA proteins. The latter proteins, along with beta-barrel protein YfaZ and VirK/YbjX family proteins, were the top overexpressed cell envelope proteins. Furthermore, the l-arginine biosynthesis pathway and putrescine-ornithine antiporter PotE were downregulated in both transcriptomes. In contrast, the expression of two pyruvate transporters (YhjX and YjiY) and genes involved in pyruvate metabolism, as well as genes involved in generating proton motive force (PMF), was antimicrobial specific. Despite the similarity of the cell envelope transcriptomes, distinctly remodeled carbon metabolism (i.e., toward fermentation of pyruvate to acetoin [colistin] and to the glyoxylate pathway [CSA-13]) distinguished both antimicrobials, which possibly reflects the intensity of the stress exerted by both agents. IMPORTANCE Colistin and ceragenins, like CSA-13, are cationic antimicrobials that disrupt the bacterial cell envelope through different mechanisms. Here, we examined the genomic and transcriptome changes in Enterobacter hormaechei ST89, an emerging hospital pathogen, after prolonged exposure to these agents to identify potential resistance mechanisms. Interestingly, we observed downregulation of genes associated with acid stress response as well as distinct dysregulation of genes involved in carbon metabolism, resulting in a switch from pyruvate fermentation to acetoin (colistin) and the glyoxylate pathway (CSA-13). Therefore, we hypothesize that repression of the acid stress response, which alkalinizes cytoplasmic pH and, in turn, suppresses resistance to cationic antimicrobials, could be interpreted as an adaptation that prevents alkalinization of cytoplasmic pH in emergencies induced by colistin and CSA-13. Consequently, this alteration critical for cell physiology must be compensated via remodeling carbon and/or amino acid metabolism to limit acidic by-product production.
Keywords: CAMPs; CSA-13; RNA-seq; adaptive resistance; carbon metabolism; ceragenins; colistin; next-generation sequencing; stress response; transcriptional regulation.
Conflict of interest statement
The authors declare a conflict of interest. Robert Bucki received funding from N8 Medical, which focuses on commercialization of medical devices that incorporate ceragenins, to attend TechnoSepsis meeting on Oct 5-7, 2022 in Spain.
Figures
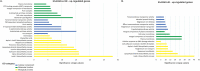



Similar articles
-
MgrB Inactivation Is Responsible for Acquired Resistance to Colistin in Enterobacter hormaechei subsp. steigerwaltii.Antimicrob Agents Chemother. 2020 May 21;64(6):e00128-20. doi: 10.1128/AAC.00128-20. Print 2020 May 21. Antimicrob Agents Chemother. 2020. PMID: 32253218 Free PMC article.
-
Ceragenins and Antimicrobial Peptides Kill Bacteria through Distinct Mechanisms.mBio. 2022 Feb 22;13(1):e0272621. doi: 10.1128/mbio.02726-21. Epub 2022 Jan 25. mBio. 2022. PMID: 35073755 Free PMC article.
-
CSA-131, a ceragenin active against colistin-resistant Acinetobacter baumannii and Pseudomonas aeruginosa clinical isolates.Int J Antimicrob Agents. 2015 Nov;46(5):568-71. doi: 10.1016/j.ijantimicag.2015.08.003. Epub 2015 Sep 7. Int J Antimicrob Agents. 2015. PMID: 26395218
-
Antibiotic resistance in Enterobacter hormaechei.Int J Antimicrob Agents. 2022 Oct;60(4):106650. doi: 10.1016/j.ijantimicag.2022.106650. Epub 2022 Aug 4. Int J Antimicrob Agents. 2022. PMID: 35934231 Review.
-
Colistin and tigecycline resistance in carbapenemase-producing Gram-negative bacteria: emerging resistance mechanisms and detection methods.J Appl Microbiol. 2016 Sep;121(3):601-17. doi: 10.1111/jam.13169. Epub 2016 Jul 4. J Appl Microbiol. 2016. PMID: 27153928 Review.
Cited by
-
Survival and adaptative strategies of Enterotoxigenic E. coli (ETEC) to the freshwater environment.Res Sq [Preprint]. 2025 Mar 19:rs.3.rs-6252921. doi: 10.21203/rs.3.rs-6252921/v1. Res Sq. 2025. PMID: 40166005 Free PMC article. Preprint.
-
Natural Antimicrobial Peptides and Their Synthetic Analogues for Effective Oral Microflora Control and Oral Infection Treatment-The Role of Ceragenins in the Development of New Therapeutic Methods.Pharmaceuticals (Basel). 2024 Dec 20;17(12):1725. doi: 10.3390/ph17121725. Pharmaceuticals (Basel). 2024. PMID: 39770567 Free PMC article. Review.
References
-
- Matuschek E, Brolund A, Karlsson Lindsjö O, Giske CG, Byfors S, Kahlmeter G. 2021. Revisiting colistin susceptibility testing: will adding calcium to Mueller-Hinton agar improve the detection of colistin resistance? Clin Microbiol Infect 27:1172.e1–1172.e5. doi:10.1016/j.cmi.2021.04.014. - DOI - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources

