A critical region of A20 unveiled by missense TNFAIP3 variations that lead to autoinflammation
- PMID: 37342083
- PMCID: PMC10284599
- DOI: 10.7554/eLife.81280
A critical region of A20 unveiled by missense TNFAIP3 variations that lead to autoinflammation
Abstract
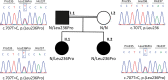
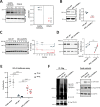
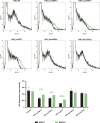
A20 haploinsufficiency (HA20) is an autoinflammatory disease caused by heterozygous loss-of-function variations in TNFAIP3, the gene encoding the A20 protein. Diagnosis of HA20 is challenging due to its heterogeneous clinical presentation and the lack of pathognomonic symptoms. While the pathogenic effect of TNFAIP3 truncating variations is clearly established, that of missense variations is difficult to determine. Herein, we identified a novel TNFAIP3 variation, p.(Leu236Pro), located in the A20 ovarian tumor (OTU) domain and demonstrated its pathogenicity. In the patients' primary cells, we observed reduced A20 levels. Protein destabilization was predicted in silico for A20_Leu236Pro and enhanced proteasomal degradation was confirmed in vitro through a flow cytometry-based functional assay. By applying this approach to the study of another missense variant, A20_Leu275Pro, for which no functional characterization has been performed to date, we showed that this variant also undergoes enhanced proteasomal degradation. Moreover, we showed a disrupted ability of A20_Leu236Pro to inhibit the NF-κB pathway and to deubiquitinate its substrate TRAF6. Structural modeling revealed that two residues involved in OTU pathogenic missense variations (i.e. Glu192Lys and Cys243Tyr) establish common interactions with Leu236. Interpretation of newly identified missense variations is challenging, requiring, as illustrated here, functional demonstration of their pathogenicity. Together with functional studies, in silico structure analysis is a valuable approach that allowed us (i) to provide a mechanistic explanation for the haploinsufficiency resulting from missense variations and (ii) to unveil a region within the OTU domain critical for A20 function.
Keywords: A20; A20 haploinsufficiency; TNFAIP3; genetics; genomics; human; immunology; inflammation; missense variation; pathogenic significance; proteasomal degradation.
© 2023, El Khouri et al.
Conflict of interest statement
EE, FD, CL, EA, AD, AN, WP, SD, AD, BC, FD, SK, SA, AA, IG No competing interests declared
Figures



Update of
- doi: 10.1101/2022.07.29.502017
References
-
- Aslani N, Asnaashari K, Parvaneh N, Shahrooei M, Sotoudeh-Anvari M, Shahram F, Ziaee V. Tnfaip3 Mutation causing Haploinsufficiency of A20 with a Hemophagocytic Lymphohistiocytosis phenotype: a report of two cases. Pediatric Rheumatology Online Journal. 2022;20:78. doi: 10.1186/s12969-022-00735-1. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

