Architecture of the cortical actomyosin network driving apical constriction in C. elegans
- PMID: 37351566
- PMCID: PMC10289891
- DOI: 10.1083/jcb.202302102
Architecture of the cortical actomyosin network driving apical constriction in C. elegans
Abstract
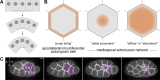
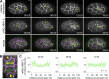
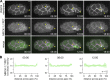
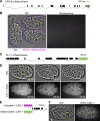
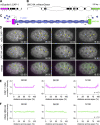
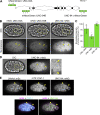
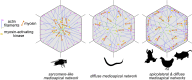
Apical constriction is a cell shape change that drives key morphogenetic events during development, including gastrulation and neural tube formation. The forces driving apical constriction are primarily generated through the contraction of apicolateral and/or medioapical actomyosin networks. In the Drosophila ventral furrow, the medioapical actomyosin network has a sarcomere-like architecture, with radially polarized actin filaments and centrally enriched non-muscle myosin II and myosin activating kinase. To determine if this is a broadly conserved actin architecture driving apical constriction, we examined actomyosin architecture during C. elegans gastrulation, in which two endodermal precursor cells internalize from the surface of the embryo. Quantification of protein localization showed that neither the non-muscle myosin II NMY-2 nor the myosin-activating kinase MRCK-1 is enriched at the center of the apex. Further, visualization of barbed- and pointed-end capping proteins revealed that actin filaments do not exhibit radial polarization at the apex. Our results demonstrate that C. elegans endodermal precursor cells apically constrict using a mixed-polarity actin filament network and with myosin and a myosin activator distributed throughout the network. Taken together with observations made in other organisms, our results demonstrate that diverse actomyosin architectures are used in animal cells to accomplish apical constriction.
© 2023 Zhang et al.
Conflict of interest statement
Disclosures: The authors declare no competing interests exist.
Figures

Update of
-
Architecture of the cortical actomyosin network driving apical constriction in C. elegans.bioRxiv [Preprint]. 2023 Feb 1:2023.01.30.526280. doi: 10.1101/2023.01.30.526280. bioRxiv. 2023. Update in: J Cell Biol. 2023 Sep 4;222(9):e202302102. doi: 10.1083/jcb.202302102. PMID: 36778218 Free PMC article. Updated. Preprint.
Similar articles
-
Architecture of the cortical actomyosin network driving apical constriction in C. elegans.bioRxiv [Preprint]. 2023 Feb 1:2023.01.30.526280. doi: 10.1101/2023.01.30.526280. bioRxiv. 2023. Update in: J Cell Biol. 2023 Sep 4;222(9):e202302102. doi: 10.1083/jcb.202302102. PMID: 36778218 Free PMC article. Updated. Preprint.
-
MRCK-1 Drives Apical Constriction in C. elegans by Linking Developmental Patterning to Force Generation.Curr Biol. 2016 Aug 22;26(16):2079-89. doi: 10.1016/j.cub.2016.06.010. Epub 2016 Jul 21. Curr Biol. 2016. PMID: 27451898 Free PMC article.
-
Apical Sarcomere-like Actomyosin Contracts Nonmuscle Drosophila Epithelial Cells.Dev Cell. 2016 Nov 7;39(3):346-358. doi: 10.1016/j.devcel.2016.09.023. Epub 2016 Oct 20. Dev Cell. 2016. PMID: 27773487 Free PMC article.
-
Roles of Actin in the Morphogenesis of the Early Caenorhabditis elegans Embryo.Int J Mol Sci. 2020 May 21;21(10):3652. doi: 10.3390/ijms21103652. Int J Mol Sci. 2020. PMID: 32455793 Free PMC article. Review.
-
Caenorhabditis elegans Gastrulation: A Model for Understanding How Cells Polarize, Change Shape, and Journey Toward the Center of an Embryo.Genetics. 2020 Feb;214(2):265-277. doi: 10.1534/genetics.119.300240. Genetics. 2020. PMID: 32029580 Free PMC article. Review.
Cited by
-
Cell signaling facilitates apical constriction by basolaterally recruiting Arp2/3 via Rac and WAVE.bioRxiv [Preprint]. 2024 Sep 23:2024.09.23.614059. doi: 10.1101/2024.09.23.614059. bioRxiv. 2024. Update in: J Cell Biol. 2025 May 05;224(5):e202409133. doi: 10.1083/jcb.202409133. PMID: 39386716 Free PMC article. Updated. Preprint.
-
Cell signaling facilitates apical constriction by basolaterally recruiting Arp2/3 via Rac and WAVE.J Cell Biol. 2025 May 5;224(5):e202409133. doi: 10.1083/jcb.202409133. Epub 2025 Mar 5. J Cell Biol. 2025. PMID: 40042443
References
-
- Burnside, B. 1973. Microtubules and microfilaments in amphibian neurulation. Am. Zool. 13:989–1006. 10.1093/icb/13.4.989 - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

