Analysis of Genome-Wide Mutational Dependence in Naturally Evolving Mycobacterium tuberculosis Populations
- PMID: 37352142
- PMCID: PMC10292908
- DOI: 10.1093/molbev/msad131
Analysis of Genome-Wide Mutational Dependence in Naturally Evolving Mycobacterium tuberculosis Populations
Abstract
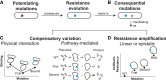
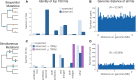
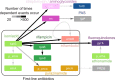
Pathogenic microorganisms are in a perpetual struggle for survival in changing host environments, where host pressures necessitate changes in pathogen virulence, antibiotic resistance, or transmissibility. The genetic basis of phenotypic adaptation by pathogens is difficult to study in vivo. In this work, we develop a phylogenetic method to detect genetic dependencies that promote pathogen adaptation using 31,428 in vivo sampled Mycobacterium tuberculosis genomes, a globally prevalent bacterial pathogen with increasing levels of antibiotic resistance. We find that dependencies between mutations are enriched in antigenic and antibiotic resistance functions and discover 23 mutations that potentiate the development of antibiotic resistance. Between 11% and 92% of resistant strains harbor a dependent mutation acquired after a resistance-conferring variant. We demonstrate the pervasiveness of genetic dependency in adaptation of naturally evolving populations and the utility of the proposed computational approach.
Keywords: computational biology; microbial evolution; microbial genomics; microbiology; mycobacterium.
© The Author(s) 2023. Published by Oxford University Press on behalf of Society for Molecular Biology and Evolution.
Figures


References
-
- Allen AC, Malaga W, Gaudin C, Volle A, Moreau F, Hassan A, Astarie-Dequeker C, Peixoto A, Antoine R, Pawlik A, et al. . 2021. Parallel in vivo experimental evolution reveals that increased stress resistance was key for the emergence of persistent tuberculosis bacilli. Nat Microbiol. 6:1082–1093. - PubMed
-
- Andersson DI, Hughes D. 2010. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat Rev Microbiol. 8:260–271. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

