An unnatural enzyme with endonuclease activity towards small non-coding RNAs
- PMID: 37355703
- PMCID: PMC10290691
- DOI: 10.1038/s41467-023-39105-0
An unnatural enzyme with endonuclease activity towards small non-coding RNAs
Erratum in
-
Author Correction: An unnatural enzyme with endonuclease activity towards small non-coding RNAs.Nat Commun. 2023 Nov 21;14(1):7588. doi: 10.1038/s41467-023-43521-7. Nat Commun. 2023. PMID: 37990004 Free PMC article. No abstract available.
Abstract
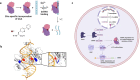
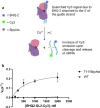
Endonucleases are enzymes that cleave internal phosphodiester bonds within double-stranded DNA or RNA and are essential for biological functions. Herein, we use genetic code expansion to create an unnatural endonuclease that cleaves non-coding RNAs including short interfering RNA (siRNA) and microRNAs (miRNAs), a function that does not exist in nature. We introduce a metal-chelating unnatural amino acid, (2,2'-bipyridin-5-yl)alanine (BpyAla) to impart endonuclease activity to the viral suppressor of RNA silencing protein p19. Upon binding of copper, the mutant p19-T111BpyAla displays catalytic site-specific cleavage of siRNA and human miRNAs. Catalysis is confirmed using fluorescence polarization and fluorescence turn-on. Global miRNA profiling reveals that the engineered enzyme cleaves miRNAs in a human cell line. The therapeutic potential is demonstrated by targeting miR-122, a critical host factor for the hepatitis C virus (HCV). Unnatural endonuclease function is shown to deplete miR-122 levels with similar effects to an antagomir that reduces HCV levels therapeutically.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

