Extracting nanoscale membrane morphology from single-molecule localizations
- PMID: 37355772
- PMCID: PMC10432223
- DOI: 10.1016/j.bpj.2023.06.010
Extracting nanoscale membrane morphology from single-molecule localizations
Abstract
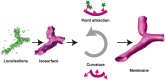
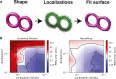
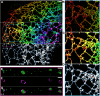
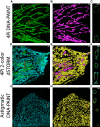
Membrane surface reconstruction at the nanometer scale is required for understanding mechanisms of subcellular shape change. This historically has been the domain of electron microscopy, but extraction of surfaces from specific labels is a difficult task in this imaging modality. Existing methods for extracting surfaces from fluorescence microscopy have poor resolution or require high-quality super-resolution data that are manually cleaned and curated. Here, we present NanoWrap, a new method for extracting surfaces from generalized single-molecule localization microscopy data. This makes it possible to study the shape of specifically labeled membranous structures inside cells. We validate NanoWrap using simulations and demonstrate its reconstruction capabilities on single-molecule localization microscopy data of the endoplasmic reticulum and mitochondria. NanoWrap is implemented in the open-source Python Microscopy Environment.
Copyright © 2023 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests J.B. discloses a significant financial interest in Bruker Corp., Hamamatsu Photonics, and panluminate Inc.
Figures
Update of
-
Extracting nanoscale membrane morphology from single-molecule localizations.bioRxiv [Preprint]. 2023 Jun 17:2023.01.26.525798. doi: 10.1101/2023.01.26.525798. bioRxiv. 2023. Update in: Biophys J. 2023 Aug 8;122(15):3022-3030. doi: 10.1016/j.bpj.2023.06.010. PMID: 36945449 Free PMC article. Updated. Preprint.
References
-
- Westrate L.M., Lee J.E., et al. Voeltz G.K. Form Follows Function: The Importance of Endoplasmic Reticulum Shape. Annu. Rev. Biochem. 2015;84:791–811. - PubMed
-
- Kaksonen M., Roux A. Mechanisms of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 2018;19:313–326. http://www.nature.com/articles/nrm.2017.132 - PubMed
-
- Haupt A., Minc N. How cells sense their own shape – mechanisms to probe cell geometry and their implications in cellular organization and function. J. Cell Sci. 2018;131:jcs214015. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

