CsrA Positively and Directly Regulates the Expression of the pdu, pocR, and eut Genes Required for the Luminal Replication of Salmonella Typhimurium
- PMID: 37358421
- PMCID: PMC10433801
- DOI: 10.1128/spectrum.01516-23
CsrA Positively and Directly Regulates the Expression of the pdu, pocR, and eut Genes Required for the Luminal Replication of Salmonella Typhimurium
Abstract
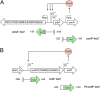
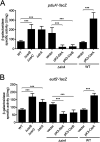
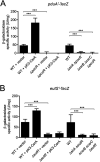
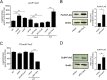
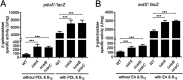
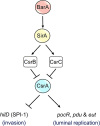
Enteric pathogens, such as Salmonella, have evolved to thrive in the inflamed gut. Genes located within the Salmonella pathogenicity island 1 (SPI-1) mediate the invasion of cells from the intestinal epithelium and the induction of an intestinal inflammatory response. Alternative electron acceptors become available in the inflamed gut and are utilized by Salmonella for luminal replication through the metabolism of propanediol and ethanolamine, using the enzymes encoded by the pdu and eut genes. The RNA-binding protein CsrA inhibits the expression of HilD, which is the central transcriptional regulator of the SPI-1 genes. Previous studies suggest that CsrA also regulates the expression of the pdu and eut genes, but the mechanism for this regulation is unknown. In this work, we show that CsrA positively regulates the pdu genes by binding to the pocR and pduA transcripts as well as the eut genes by binding to the eutS transcript. Furthermore, our results show that the SirA-CsrB/CsrC-CsrA regulatory cascade controls the expression of the pdu and eut genes mediated by PocR or EutR, which are the positive AraC-like transcriptional regulators for the pdu and eut genes, respectively. By oppositely regulating the expression of genes for invasion and for luminal replication, the SirA-CsrB/CsrC-CsrA regulatory cascade could be involved in the generation of two Salmonella populations that cooperate for intestinal colonization and transmission. Our study provides new insight into the regulatory mechanisms that govern Salmonella virulence. IMPORTANCE The regulatory mechanisms that control the expression of virulence genes are essential for bacteria to infect hosts. Salmonella has developed diverse regulatory mechanisms to colonize the host gut. For instance, the SirA-CsrB/CsrC-CsrA regulatory cascade controls the expression of the SPI-1 genes, which are required for this bacterium to invade intestinal epithelium cells and for the induction of an intestinal inflammatory response. In this study, we determine the mechanisms by which the SirA-CsrB/CsrC-CsrA regulatory cascade controls the expression of the pdu and eut genes, which are necessary for the replication of Salmonella in the intestinal lumen. Thus, our data, together with the results of previous reports, indicate that the SirA-CsrB/CsrC-CsrA regulatory cascade has an important role in the intestinal colonization by Salmonella.
Keywords: Csr; Salmonella; regulation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
SirA, CsrBC and HilD form in vivo a regulatory cascade that controls the SP1-1 and SPI-2 gene expression when Salmonella Typhimurium is in the intestinal lumen and are required for cecal colonization and liver dissemination in the avian model.Arch Microbiol. 2025 Apr 2;207(5):108. doi: 10.1007/s00203-025-04305-3. Arch Microbiol. 2025. PMID: 40169403 Free PMC article.
-
Proteomic analysis reveals the global effect of the BarA/SirA-Csr regulatory cascade in Salmonella Typhimurium grown in conditions that favor the expression of invasion genes.J Proteomics. 2023 Aug 30;286:104960. doi: 10.1016/j.jprot.2023.104960. Epub 2023 Jul 13. J Proteomics. 2023. PMID: 37451358
-
An incoherent feedforward loop formed by SirA/BarA, HilE and HilD is involved in controlling the growth cost of virulence factor expression by Salmonella Typhimurium.PLoS Pathog. 2021 May 28;17(5):e1009630. doi: 10.1371/journal.ppat.1009630. eCollection 2021 May. PLoS Pathog. 2021. PMID: 34048498 Free PMC article.
-
Global regulation by the small RNA-binding protein CsrA and the non-coding RNA molecule CsrB.Mol Microbiol. 1998 Sep;29(6):1321-30. doi: 10.1046/j.1365-2958.1998.01021.x. Mol Microbiol. 1998. PMID: 9781871 Review.
-
Prokaryotic Organelles: Bacterial Microcompartments in E. coli and Salmonella.EcoSal Plus. 2020 Oct;9(1):10.1128/ecosalplus.ESP-0025-2019. doi: 10.1128/ecosalplus.ESP-0025-2019. EcoSal Plus. 2020. PMID: 33030141 Free PMC article. Review.
Cited by
-
SirA, CsrBC and HilD form in vivo a regulatory cascade that controls the SP1-1 and SPI-2 gene expression when Salmonella Typhimurium is in the intestinal lumen and are required for cecal colonization and liver dissemination in the avian model.Arch Microbiol. 2025 Apr 2;207(5):108. doi: 10.1007/s00203-025-04305-3. Arch Microbiol. 2025. PMID: 40169403 Free PMC article.
-
Phage-mediated horizontal transfer of Salmonella enterica virulence genes with regulatory feedback from the host.Imeta. 2025 May 20;4(4):e70042. doi: 10.1002/imt2.70042. eCollection 2025 Aug. Imeta. 2025. PMID: 40860440 Free PMC article.
-
Nanoplastics-mediated physiologic and genomic responses in pathogenic Escherichia coli O157:H7.J Nanobiotechnology. 2025 Apr 21;23(1):304. doi: 10.1186/s12951-025-03369-z. J Nanobiotechnology. 2025. PMID: 40259296 Free PMC article.
-
Post-Transcriptional Regulation of the MiaA Prenyl Transferase by CsrA and the Small RNA CsrB in Escherichia coli.Int J Mol Sci. 2025 Jun 24;26(13):6068. doi: 10.3390/ijms26136068. Int J Mol Sci. 2025. PMID: 40649849 Free PMC article.
References
-
- Price-Carter M, Tingey J, Bobik TA, Roth JR. 2001. The alternative electron acceptor tetrathionate supports B12-dependent anaerobic growth of Salmonella enterica serovar Typhimurium on ethanolamine or 1,2-propanediol. J Bacteriol 183:2463–2475. doi:10.1128/JB.183.8.2463-2475.2001. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

