Genomic and Functional Characterization of Longitudinal Pseudomonas aeruginosa Isolates from Young Patients with Cystic Fibrosis
- PMID: 37358436
- PMCID: PMC10433850
- DOI: 10.1128/spectrum.01556-23
Genomic and Functional Characterization of Longitudinal Pseudomonas aeruginosa Isolates from Young Patients with Cystic Fibrosis
Abstract
Individuals with cystic fibrosis (CF) suffer from frequent and recurring microbial airway infections. The Gram-negative bacterium Pseudomonas aeruginosa is one of the most common organisms isolated from CF patient airways. P. aeruginosa establishes chronic infections that persist throughout a patient's lifetime and is a major cause of morbidity and mortality. Throughout the course of infection, P. aeruginosa must evolve and adapt from an initial state of early, transient colonization to chronic colonization of the airways. Here, we examined isolates of P. aeruginosa from children under the age of 3 years old with CF to determine genetic adaptations the bacterium undergoes during this early stage of colonization and infection. These isolates were collected when early aggressive antimicrobial therapy was not the standard of care and therefore highlight strain evolution under limited antibiotic pressure. Examination of specific phenotypic adaptations, such as lipid A palmitoylation, antibiotic resistance, and loss of quorum sensing, did not reveal a clear genetic basis for such changes. Additionally, we demonstrate that the geography of patient origin, within the United States or among other countries, does not appear to significantly influence genetic adaptation. In summary, our results support the long-standing model that patients acquire individual isolates of P. aeruginosa that subsequently become hyperadapted to the patient-specific airway environment. This study provides a multipatient genomic analysis of isolates from young CF patients in the United States and contributes data regarding early colonization and adaptation to the growing body of research about P. aeruginosa evolution in the context of CF airway disease. IMPORTANCE Chronic lung infection with Pseudomonas aeruginosa is of major concern for patients with cystic fibrosis (CF). During infection, P. aeruginosa undergoes genomic and functional adaptation to the hyperinflammatory CF airway, resulting in worsening lung function and pulmonary decline. All studies that describe these adaptations use P. aeruginosa obtained from older children or adults during late chronic lung infection; however, children with CF can be infected with P. aeruginosa as early as 3 months of age. Therefore, it is unclear when these genomic and functional adaptations occur over the course of CF lung infection, as access to P. aeruginosa isolates in children during early infection is limited. Here, we present a unique cohort of CF patients who were identified as being infected with P. aeruginosa at an early age prior to aggressive antibiotic therapy. Furthermore, we performed genomic and functional characterization of these isolates to address whether chronic CF P. aeruginosa phenotypes are present during early infection.
Keywords: LPS evolution; Pseudomonas aeruginosa; airway adaptation; cystic fibrosis; genomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

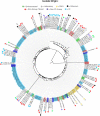
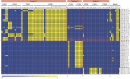
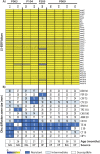
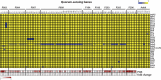
Similar articles
-
Pseudomonas aeruginosa Volatilome Characteristics and Adaptations in Chronic Cystic Fibrosis Lung Infections.mSphere. 2020 Oct 7;5(5):e00843-20. doi: 10.1128/mSphere.00843-20. mSphere. 2020. PMID: 33028687 Free PMC article.
-
Characterization of Pseudomonas aeruginosa from subjects with diffuse panbronchiolitis.Microbiol Spectr. 2024 Nov 5;12(11):e0053024. doi: 10.1128/spectrum.00530-24. Epub 2024 Oct 8. Microbiol Spectr. 2024. PMID: 39377602 Free PMC article.
-
Genotypic and Phenotypic Diversity of Staphylococcus aureus Isolates from Cystic Fibrosis Patient Lung Infections and Their Interactions with Pseudomonas aeruginosa.mBio. 2020 Jun 23;11(3):e00735-20. doi: 10.1128/mBio.00735-20. mBio. 2020. PMID: 32576671 Free PMC article.
-
Microevolution of Pseudomonas aeruginosa to a chronic pathogen of the cystic fibrosis lung.Curr Top Microbiol Immunol. 2013;358:91-118. doi: 10.1007/82_2011_199. Curr Top Microbiol Immunol. 2013. PMID: 22311171 Review.
-
Adaptation of Pseudomonas aeruginosa to the cystic fibrosis airway: an evolutionary perspective.Nat Rev Microbiol. 2012 Dec;10(12):841-51. doi: 10.1038/nrmicro2907. Epub 2012 Nov 13. Nat Rev Microbiol. 2012. PMID: 23147702 Review.
Cited by
-
Whole Genome Analysis of Ocular Pseudomonas aeruginosa Isolates Reveals Genetic Diversity.Invest Ophthalmol Vis Sci. 2025 Jun 2;66(6):58. doi: 10.1167/iovs.66.6.58. Invest Ophthalmol Vis Sci. 2025. PMID: 40530921 Free PMC article.
-
Pseudomonas aeruginosa Lipid A Structural Variants Induce Altered Immune Responses.Am J Respir Cell Mol Biol. 2024 Aug;71(2):207-218. doi: 10.1165/rcmb.2024-0059OC. Am J Respir Cell Mol Biol. 2024. PMID: 38656811 Free PMC article.
-
Divergent Pseudomonas aeruginosa LpxO enzymes perform site-specific lipid A 2-hydroxylation.mBio. 2024 Feb 14;15(2):e0282323. doi: 10.1128/mbio.02823-23. Epub 2023 Dec 22. mBio. 2024. PMID: 38131669 Free PMC article.
-
Resistance of Pseudomonas aeruginosa to Antibiotics During Long-Term Persistence in Patients with Cystic Fibrosis.Antibiotics (Basel). 2025 Mar 14;14(3):302. doi: 10.3390/antibiotics14030302. Antibiotics (Basel). 2025. PMID: 40149112 Free PMC article.
-
Biofilm Formation of Pseudomonas aeruginosa in Cystic Fibrosis: Mechanisms of Persistence, Adaptation, and Pathogenesis.Microorganisms. 2025 Jun 30;13(7):1527. doi: 10.3390/microorganisms13071527. Microorganisms. 2025. PMID: 40732035 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

