Prognostic and immune infiltration significance of ARID1A in TCGA molecular subtypes of gastric adenocarcinoma
- PMID: 37366273
- PMCID: PMC10501255
- DOI: 10.1002/cam4.6294
Prognostic and immune infiltration significance of ARID1A in TCGA molecular subtypes of gastric adenocarcinoma
Abstract
Background: AT-rich interaction domain 1A (ARID1A) is an essential subunit of the switch/sucrose non-fermentable chromatin remodeling complex and is considered to be a tumor suppressor. The Cancer Genome Atlas (TCGA) molecular classification has deepened our understanding of gastric cancer at the molecular level. This study explored the significance of ARID1A expression in TCGA subtypes of gastric adenocarcinoma.
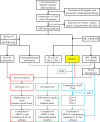
Methods: We collected 1248 postoperative patients with gastric adenocarcinoma, constructed tissue microarrays, performed immunohistochemistry for ARID1A, and obtained correlations between ARID1A and clinicopathological variables. We then carried out the prognostic analysis of ARID1A in TCGA subtypes. Finally, we screened patients by random sampling and propensity score matching method and performed multiplex immunofluorescence to explore the effects of ARID1A on CD4, CD8, and PD-L1 expression in TCGA subtypes.
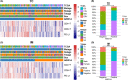
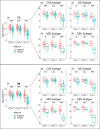
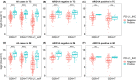
Results: Seven variables independently associated with ARID1A were screened out: mismatch repair proteins, PD-L1, T stage, differentiation status, p53, E-cadherin, and EBER. The independent prognostic variables in the genomically stable (GS) subtype were N stage, M stage, T stage, chemotherapy, size, and ARID1A. PD-L1 expression was higher in the ARID1A negative group than in the ARID1A positive group in all TCGA subgroups. CD4 showed higher expression in the ARID1A negative group in most subtypes, while CD8 did not show the difference in most subtypes. When ARID1A was negative, PD-L1 expression was positively correlated with CD4/CD8 expression; while when ARID1A was positive, this correlation disappeared.
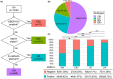
Conclusions: The negative expression of ARID1A occurred more frequently in the Epstein-Barr virus and microsatellite instability subtypes and was an independent adverse prognostic factor in the GS subtype. In the TCGA subtypes, ARID1A negative expression caused increased CD4 and PD-L1 expression, whereas CD8 expression appeared independent of ARID1A. The expression of CD4/CD8 induced by ARID1A negativity was accompanied by an increase in PD-L1 expression.
Keywords: ARID1A; TCGA subtype; gastric adenocarcinoma; immune infiltration; prognosis.
© 2023 The Authors. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures






References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

