Development of a Redox-Polymer-Based Electrochemical Glucose Biosensor Suitable for Integration in Microfluidic 3D Cell Culture Systems
- PMID: 37366947
- PMCID: PMC10295976
- DOI: 10.3390/bios13060582
Development of a Redox-Polymer-Based Electrochemical Glucose Biosensor Suitable for Integration in Microfluidic 3D Cell Culture Systems
Abstract
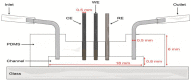
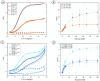
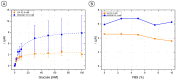
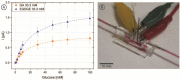
The inclusion of online, in situ biosensors in microfluidic cell cultures is important to monitor and characterize a physiologically mimicking environment. This work presents the performance of second-generation electrochemical enzymatic biosensors to detect glucose in cell culture media. Glutaraldehyde and ethylene glycol diglycidyl ether (EGDGE) were tested as cross-linkers to immobilize glucose oxidase and an osmium-modified redox polymer on the surface of carbon electrodes. Tests employing screen printed electrodes showed adequate performance in a Roswell Park Memorial Institute (RPMI-1640) media spiked with fetal bovine serum (FBS). Comparable first-generation sensors were shown to be heavily affected by complex biological media. This difference is explained in terms of the respective charge transfer mechanisms. Under the tested conditions, electron hopping between Os redox centers was less vulnerable than H2O2 diffusion to biofouling by the substances present in the cell culture matrix. By employing pencil leads as electrodes, the incorporation of these electrodes in a polydimethylsiloxane (PDMS) microfluidic channel was achieved simply and at a low cost. Under flow conditions, electrodes fabricated using EGDGE presented the best performance with a limit of detection of 0.5 mM, a linear range up to 10 mM, and a sensitivity of 4.69 μA mM-1 cm-2.
Keywords: 3D cell culture; EGDGE; cross-linking; glucose electrochemical biosensor; glucose oxidase; glutaraldehyde; on-chip evaluation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Microfluidic Electrochemical Glucose Biosensor with In Situ Enzyme Immobilization.Biosensors (Basel). 2023 Mar 9;13(3):364. doi: 10.3390/bios13030364. Biosensors (Basel). 2023. PMID: 36979576 Free PMC article.
-
A novel 3D paper-based microfluidic electrochemical glucose biosensor based on rGO-TEPA/PB sensitive film.Anal Chim Acta. 2020 Feb 1;1096:34-43. doi: 10.1016/j.aca.2019.10.049. Epub 2019 Oct 23. Anal Chim Acta. 2020. PMID: 31883589
-
Enzyme Immobilization and Mediation with Osmium Redox Polymers.Methods Mol Biol. 2017;1504:165-179. doi: 10.1007/978-1-4939-6499-4_13. Methods Mol Biol. 2017. PMID: 27770421
-
Enzyme-based amperometric galactose biosensors: a review.Mikrochim Acta. 2017;184(10):3663-3671. doi: 10.1007/s00604-017-2465-z. Epub 2017 Aug 25. Mikrochim Acta. 2017. PMID: 28979051 Free PMC article. Review.
-
Polymers and Plastics Modified Electrodes for Biosensors: A Review.Molecules. 2020 May 24;25(10):2446. doi: 10.3390/molecules25102446. Molecules. 2020. PMID: 32456314 Free PMC article. Review.
Cited by
-
Probing the influence of crosslinkers on the properties, response, and degradation of enzymatic hydrogels for electrochemical glucose biosensing through fluorescence analysis.RSC Adv. 2024 Mar 20;14(14):9514-9528. doi: 10.1039/d4ra00265b. eCollection 2024 Mar 20. RSC Adv. 2024. PMID: 38516160 Free PMC article.
-
Whole Cells of Microorganisms-A Powerful Bioanalytical Tool for Measuring Integral Parameters of Pollution: A Review.Biosensors (Basel). 2025 May 4;15(5):290. doi: 10.3390/bios15050290. Biosensors (Basel). 2025. PMID: 40422029 Free PMC article. Review.
-
Recent Advances in Amperometric Biosensors for Medical Applications: A Mini-Review.Curr Top Med Chem. 2025;25(7):741-760. doi: 10.2174/0115680266323004241015122441. Curr Top Med Chem. 2025. PMID: 39473247 Review.
References
-
- Ahmed T. Organ-on-a-chip microengineering for bio-mimicking disease models and revolutionizing drug discovery. Biosens. Bioelectron. X. 2022;11:100194. doi: 10.1016/j.biosx.2022.100194. - DOI
-
- Santbergen M.J.C., van der Zande M., Bouwmeester H., Nielen M.W.F. Online and in situ analysis of organs-on-a-chip. TrAC Trends Anal. Chem. 2019;115:138–146. doi: 10.1016/j.trac.2019.04.006. - DOI
-
- Bavli D., Prill S., Ezra E., Levy G., Cohen M., Vinken M., Vanfleteren J., Jaeger M., Nahmias Y. Real-time monitoring of metabolic function in liver-onchip microdevices tracks the dynamics of Mitochondrial dysfunction. Proc. Natl. Acad. Sci. USA. 2016;113:E2231–E2240. doi: 10.1073/pnas.1522556113. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

