Unlocking the Power of Nanopores: Recent Advances in Biosensing Applications and Analog Front-End
- PMID: 37366963
- PMCID: PMC10296294
- DOI: 10.3390/bios13060598
Unlocking the Power of Nanopores: Recent Advances in Biosensing Applications and Analog Front-End
Abstract
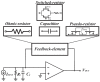
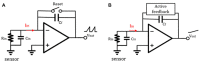
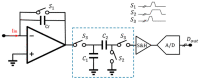
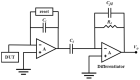
The biomedical field has always fostered innovation and the development of various new technologies. Beginning in the last century, demand for picoampere-level current detection in biomedicine has increased, leading to continuous breakthroughs in biosensor technology. Among emerging biomedical sensing technologies, nanopore sensing has shown great potential. This paper reviews nanopore sensing applications, such as chiral molecules, DNA sequencing, and protein sequencing. However, the ionic current for different molecules differs significantly, and the detection bandwidths vary as well. Therefore, this article focuses on current sensing circuits, and introduces the latest design schemes and circuit structures of different feedback components of transimpedance amplifiers mainly used in nanopore DNA sequencing.
Keywords: DNA sequencing; biosensors; chiral molecules; nanopores; protein sequencing; transimpedance amplifiers.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Solid-State Nanopores for Biomolecular Analysis and Detection.Adv Biochem Eng Biotechnol. 2024;187:283-316. doi: 10.1007/10_2023_240. Adv Biochem Eng Biotechnol. 2024. PMID: 38273209 Review.
-
Nanopore-CMOS Interfaces for DNA Sequencing.Biosensors (Basel). 2016 Aug 6;6(3):42. doi: 10.3390/bios6030042. Biosensors (Basel). 2016. PMID: 27509529 Free PMC article. Review.
-
Nanopore Technology and Its Applications in Gene Sequencing.Biosensors (Basel). 2021 Jun 30;11(7):214. doi: 10.3390/bios11070214. Biosensors (Basel). 2021. PMID: 34208844 Free PMC article. Review.
-
Nanopore sensors for single molecular protein detection: Research progress based on computer simulations.IET Nanobiotechnol. 2023 May;17(3):257-268. doi: 10.1049/nbt2.12124. Epub 2023 Mar 16. IET Nanobiotechnol. 2023. PMID: 36924083 Free PMC article. Review.
-
Theoretical and experimental studies on ionic currents in nanopore-based biosensors.IET Nanobiotechnol. 2014 Dec;8(4):247-56. doi: 10.1049/iet-nbt.2013.0017. IET Nanobiotechnol. 2014. PMID: 25429504
Cited by
-
Nanopipettes as a Potential Diagnostic Tool for Selective Nanopore Detection of Biomolecules.Biosensors (Basel). 2024 Dec 19;14(12):627. doi: 10.3390/bios14120627. Biosensors (Basel). 2024. PMID: 39727892 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

