Characterisation of Type II DNA Methyltransferases of Metamycoplasma hominis
- PMID: 37375093
- PMCID: PMC10305163
- DOI: 10.3390/microorganisms11061591
Characterisation of Type II DNA Methyltransferases of Metamycoplasma hominis
Abstract
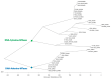
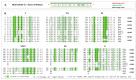
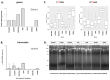
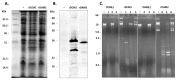
Bacterial virulence, persistence and defence are affected by epigenetic modifications, including DNA methylation. Solitary DNA methyltransferases modulate a variety of cellular processes and influence bacterial virulence; as part of a restriction-modification (RM) system, they act as a primitive immune system in methylating the own DNA, while unmethylated foreign DNA is restricted. We identified a large family of type II DNA methyltransferases in Metamycoplasma hominis, comprising six solitary methyltransferases and four RM systems. Motif-specific 5mC and 6mA methylations were identified with a tailored Tombo analysis on Nanopore reads. Selected motifs with methylation scores >0.5 fit with the gene presence of DAM1 and DAM2, DCM2, DCM3, and DCM6, but not for DCM1, whose activity was strain-dependent. The activity of DCM1 for CmCWGG and of both DAM1 and DAM2 for GmATC was proven in methylation-sensitive restriction and finally for recombinant rDCM1 and rDAM2 against a dam-, dcm-negative background. A hitherto unknown dcm8/dam3 gene fusion containing a (TA) repeat region of varying length was characterized within a single strain, suggesting the expression of DCM8/DAM3 phase variants. The combination of genetic, bioinformatics, and enzymatic approaches enabled the detection of a huge family of type II DNA MTases in M. hominis, whose involvement in virulence and defence can now be characterized in future work.
Keywords: DNA MTase; Metamycoplasma hominis; Nanopore; RM system; Tombo; dam; dcm; orphan.
Conflict of interest statement
There are no conflict of interests to declare.
Figures






References
-
- Pereyre S., Sirand-Pugnet P., Beven L., Charron A., Renaudin H., Barre A., Avenaud P., Jacob D., Couloux A., Barbe V., et al. Life on arginine for Mycoplasma hominis: Clues from its minimal genome and comparison with other human urogenital mycoplasmas. PLoS Genet. 2009;5:e1000677. doi: 10.1371/journal.pgen.1000677. - DOI - PMC - PubMed
-
- Christiansen G., Jensen L.T., Boesen T., Emmersen J., Ladefoged S.A., Schiotz L.K., Birkelund S. Molecular biology of Mycoplasma. Wien. Klin. Wochenschr. 1997;109:557–561. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

