A new framework for metabolic connectivity mapping using bolus [18F]FDG PET and kinetic modeling
- PMID: 37377103
- PMCID: PMC10676136
- DOI: 10.1177/0271678X231184365
A new framework for metabolic connectivity mapping using bolus [18F]FDG PET and kinetic modeling
Abstract
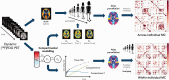
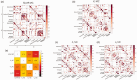
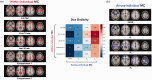
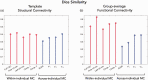
Metabolic connectivity (MC) has been previously proposed as the covariation of static [18F]FDG PET images across participants, i.e., across-individual MC (ai-MC). In few cases, MC has been inferred from dynamic [18F]FDG signals, i.e., within-individual MC (wi-MC), as for resting-state fMRI functional connectivity (FC). The validity and interpretability of both approaches is an important open issue. Here we reassess this topic, aiming to 1) develop a novel wi-MC methodology; 2) compare ai-MC maps from standardized uptake value ratio (SUVR) vs. [18F]FDG kinetic parameters fully describing the tracer behavior (i.e., Ki, K1, k3); 3) assess MC interpretability in comparison to structural connectivity and FC. We developed a new approach based on Euclidean distance to calculate wi-MC from PET time-activity curves. The across-individual correlation of SUVR, Ki, K1, k3 produced different networks depending on the chosen [18F]FDG parameter (k3 MC vs. SUVR MC, r = 0.44). We found that wi-MC and ai-MC matrices are dissimilar (maximum r = 0.37), and that the match with FC is higher for wi-MC (Dice similarity: 0.47-0.63) than for ai-MC (0.24-0.39). Our analyses demonstrate that calculating individual-level MC from dynamic PET is feasible and yields interpretable matrices that bear similarity to fMRI FC measures.
Keywords: Euclidean similarity; [18F]FDG; dynamic PET; individual-level metabolic connectivity; kinetic modeling.
Conflict of interest statement
Declaration of conflicting interestsThe author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Figures

Similar articles
-
Individual-level metabolic connectivity from dynamic [18F]FDG PET reveals glioma-induced impairments in brain architecture and offers novel insights beyond the SUVR clinical standard.Eur J Nucl Med Mol Imaging. 2025 Feb;52(3):836-850. doi: 10.1007/s00259-024-06956-8. Epub 2024 Oct 30. Eur J Nucl Med Mol Imaging. 2025. PMID: 39472368
-
Elucidating the complementarity of resting-state networks derived from dynamic [18F]FDG and hemodynamic fluctuations using simultaneous small-animal PET/MRI.Neuroimage. 2021 Aug 1;236:118045. doi: 10.1016/j.neuroimage.2021.118045. Epub 2021 Apr 10. Neuroimage. 2021. PMID: 33848625 Free PMC article.
-
Functional resting-state brain connectivity is accompanied by dynamic correlations of application-dependent [18F]FDG PET-tracer fluctuations.Neuroimage. 2019 Aug 1;196:161-172. doi: 10.1016/j.neuroimage.2019.04.034. Epub 2019 Apr 11. Neuroimage. 2019. PMID: 30981858 Free PMC article.
-
Preclinical dynamic 18F-FDG PET - tumor characterization and radiotherapy response assessment by kinetic compartment analysis.Acta Oncol. 2010 Oct;49(7):914-21. doi: 10.3109/0284186X.2010.498831. Acta Oncol. 2010. PMID: 20831478
-
Connecting the dots: approaching a standardized nomenclature for molecular connectivity in positron emission tomography.Eur J Nucl Med Mol Imaging. 2025 Jun 2. doi: 10.1007/s00259-025-07357-1. Online ahead of print. Eur J Nucl Med Mol Imaging. 2025. PMID: 40455254 Review.
Cited by
-
The brain's "dark energy" puzzle upgraded: [18F]FDG uptake, delivery and phosphorylation, and their coupling with resting-state brain activity.bioRxiv [Preprint]. 2024 Oct 7:2024.10.05.615717. doi: 10.1101/2024.10.05.615717. bioRxiv. 2024. Update in: J Cereb Blood Flow Metab. 2025 May 15:271678X251329707. doi: 10.1177/0271678X251329707. PMID: 39416159 Free PMC article. Updated. Preprint.
-
A novel method for PET connectomics guided by fibre-tracking MRI: Application to Alzheimer's disease.Hum Brain Mapp. 2024 Mar;45(4):e26659. doi: 10.1002/hbm.26659. Hum Brain Mapp. 2024. PMID: 38491564 Free PMC article.
-
Molecular connectivity studies in neurotransmission: a scoping review.Imaging Neurosci (Camb). 2025 Apr 4;3:imag_a_00530. doi: 10.1162/imag_a_00530. eCollection 2025. Imaging Neurosci (Camb). 2025. PMID: 40800788 Free PMC article.
-
The brain's "dark energy" puzzle upgraded: [18F]FDG uptake, delivery and phosphorylation, and their coupling with resting-state brain activity.J Cereb Blood Flow Metab. 2025 May 15:271678X251329707. doi: 10.1177/0271678X251329707. Online ahead of print. J Cereb Blood Flow Metab. 2025. PMID: 40370305 Free PMC article.
-
On the analysis of functional PET (fPET)-FDG: baseline mischaracterization can introduce artifactual metabolic (de)activations.bioRxiv [Preprint]. 2024 Oct 21:2024.10.17.618550. doi: 10.1101/2024.10.17.618550. bioRxiv. 2024. PMID: 39484579 Free PMC article. Preprint.
References
-
- Bertoldo A, Rizzo G, Veronese M. Deriving physiological information from PET images: from SUV to compartmental modeling. Clin Transl Imaging 2014; 2: 239–251.
-
- Betzel RF. Network neuroscience and the connectomics revolution. In: Connectomic deep brain stimulation. Elsevier: Academic Press, 2021, pp.25–58.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

