Conformation-specific Synthetic Antibodies Discriminate Multiple Functional States of the Ion Channel CorA
- PMID: 37394032
- PMCID: PMC10529903
- DOI: 10.1016/j.jmb.2023.168192
Conformation-specific Synthetic Antibodies Discriminate Multiple Functional States of the Ion Channel CorA
Abstract
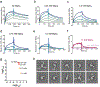
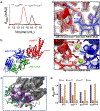
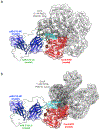
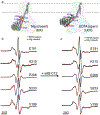
CorA, the primary magnesium ion channel in prokaryotes and archaea, is a prototypical homopentameric ion channel that undergoes ion-dependent conformational transitions. CorA adopts five-fold symmetric non-conductive states in the presence of high concentrations of Mg2+, and highly asymmetric flexible states in its complete absence. However, the latter were of insufficient resolution to be thoroughly characterized. In order to gain additional insights into the relationship between asymmetry and channel activation, we exploited phage display selection strategies to generate conformation-specific synthetic antibodies (sABs) against CorA in the absence of Mg2+. Two sABs from these selections, C12 and C18, showed different degrees of Mg2+-sensitivity. Through structural, biochemical, and biophysical characterization, we found the sABs are both conformation-specific but probe different features of the channel under open-like conditions. C18 is highly specific to the Mg2+-depleted state of CorA and through negative-stain electron microscopy (ns-EM), we show sAB binding reflects the asymmetric arrangement of CorA protomers in Mg2+-depleted conditions. We used X-ray crystallography to determine a structure at 2.0 Å resolution of sAB C12 bound to the soluble N-terminal regulatory domain of CorA. The structure shows C12 is a competitive inhibitor of regulatory magnesium binding through its interaction with the divalent cation sensing site. We subsequently exploited this relationship to capture and visualize asymmetric CorA states in different [Mg2+] using ns-EM. We additionally utilized these sABs to provide insights into the energy landscape that governs the ion-dependent conformational transitions of CorA.
Keywords: CorA; X-ray crystallography; ion channel; membrane proteins; synthetic antibodies.
Copyright © 2023 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Update of
-
Conformation-specific synthetic antibodies discriminate multiple functional states of the ion channel CorA.bioRxiv [Preprint]. 2023 May 7:2023.05.07.539746. doi: 10.1101/2023.05.07.539746. bioRxiv. 2023. Update in: J Mol Biol. 2023 Sep 1;435(17):168192. doi: 10.1016/j.jmb.2023.168192. PMID: 37205530 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

