Toward reproducible models of sequence learning: replication and analysis of a modular spiking network with reward-based learning
- PMID: 37396571
- PMCID: PMC10310927
- DOI: 10.3389/fnint.2023.935177
Toward reproducible models of sequence learning: replication and analysis of a modular spiking network with reward-based learning
Abstract
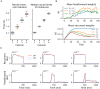
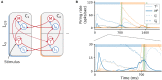
To acquire statistical regularities from the world, the brain must reliably process, and learn from, spatio-temporally structured information. Although an increasing number of computational models have attempted to explain how such sequence learning may be implemented in the neural hardware, many remain limited in functionality or lack biophysical plausibility. If we are to harvest the knowledge within these models and arrive at a deeper mechanistic understanding of sequential processing in cortical circuits, it is critical that the models and their findings are accessible, reproducible, and quantitatively comparable. Here we illustrate the importance of these aspects by providing a thorough investigation of a recently proposed sequence learning model. We re-implement the modular columnar architecture and reward-based learning rule in the open-source NEST simulator, and successfully replicate the main findings of the original study. Building on these, we perform an in-depth analysis of the model's robustness to parameter settings and underlying assumptions, highlighting its strengths and weaknesses. We demonstrate a limitation of the model consisting in the hard-wiring of the sequence order in the connectivity patterns, and suggest possible solutions. Finally, we show that the core functionality of the model is retained under more biologically-plausible constraints.
Keywords: modularity; reproducibility; reward-based learning; sequence learning model; spiking networks.
Copyright © 2023 Zajzon, Duarte and Morrison.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





References
LinkOut - more resources
Full Text Sources

