Transcriptomic response of Sinorhizobium meliloti to the predatory attack of Myxococcus xanthus
- PMID: 37405170
- PMCID: PMC10315480
- DOI: 10.3389/fmicb.2023.1213659
Transcriptomic response of Sinorhizobium meliloti to the predatory attack of Myxococcus xanthus
Abstract
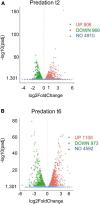
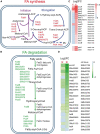
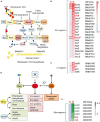
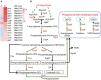
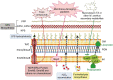
Bacterial predation impacts microbial community structures, which can have both positive and negative effects on plant and animal health and on environmental sustainability. Myxococcus xanthus is an epibiotic soil predator with a broad range of prey, including Sinorhizobium meliloti, which establishes nitrogen-fixing symbiosis with legumes. During the M. xanthus-S. meliloti interaction, the predator must adapt its transcriptome to kill and lyse the target (predatosome), and the prey must orchestrate a transcriptional response (defensome) to protect itself against the biotic stress caused by the predatory attack. Here, we describe the transcriptional changes taking place in S. meliloti in response to myxobacterial predation. The results indicate that the predator induces massive changes in the prey transcriptome with up-regulation of protein synthesis and secretion, energy generation, and fatty acid (FA) synthesis, while down-regulating genes required for FA degradation and carbohydrate transport and metabolism. The reconstruction of up-regulated pathways suggests that S. meliloti modifies the cell envelop by increasing the production of different surface polysaccharides (SPSs) and membrane lipids. Besides the barrier role of SPSs, additional mechanisms involving the activity of efflux pumps and the peptide uptake transporter BacA, together with the production of H2O2 and formaldehyde have been unveiled. Also, the induction of the iron-uptake machinery in both predator and prey reflects a strong competition for this metal. With this research we complete the characterization of the complex transcriptional changes that occur during the M. xanthus-S. meliloti interaction, which can impact the establishment of beneficial symbiosis with legumes.
Keywords: Sinorhizobium meliloti; bacterial interactions; bacterial predation; defensome; myxobacteria.
Copyright © 2023 Soto, Pérez, Muñoz-Dorado, Contreras-Moreno and Moraleda-Muñoz.
Conflict of interest statement
JP was one of the topic editors for the research topic this manuscript is submitted to: “Mechanisms of Prokaryotic Predation, Volume II.” The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Siderophores and competition for iron govern myxobacterial predation dynamics.ISME J. 2024 Jan 8;18(1):wrae077. doi: 10.1093/ismejo/wrae077. ISME J. 2024. PMID: 38696719 Free PMC article.
-
Myxococcus xanthus predation: an updated overview.Front Microbiol. 2024 Jan 24;15:1339696. doi: 10.3389/fmicb.2024.1339696. eCollection 2024. Front Microbiol. 2024. PMID: 38328431 Free PMC article. Review.
-
Copper and Melanin Play a Role in Myxococcus xanthus Predation on Sinorhizobium meliloti.Front Microbiol. 2020 Feb 4;11:94. doi: 10.3389/fmicb.2020.00094. eCollection 2020. Front Microbiol. 2020. PMID: 32117124 Free PMC article.
-
Development versus predation: Transcriptomic changes during the lifecycle of Myxococcus xanthus.Front Microbiol. 2022 Sep 26;13:1004476. doi: 10.3389/fmicb.2022.1004476. eCollection 2022. Front Microbiol. 2022. PMID: 36225384 Free PMC article.
-
The Genetics of Prey Susceptibility to Myxobacterial Predation: A Review, Including an Investigation into Pseudomonas aeruginosa Mutations Affecting Predation by Myxococcus xanthus.Microb Physiol. 2021;31(2):57-66. doi: 10.1159/000515546. Epub 2021 Apr 1. Microb Physiol. 2021. PMID: 33794538 Review.
Cited by
-
The role of microbial interactions on rhizobial fitness.Front Plant Sci. 2023 Oct 9;14:1277262. doi: 10.3389/fpls.2023.1277262. eCollection 2023. Front Plant Sci. 2023. PMID: 37877089 Free PMC article. Review.
-
Siderophores and competition for iron govern myxobacterial predation dynamics.ISME J. 2024 Jan 8;18(1):wrae077. doi: 10.1093/ismejo/wrae077. ISME J. 2024. PMID: 38696719 Free PMC article.
-
Myxococcus xanthus predation: an updated overview.Front Microbiol. 2024 Jan 24;15:1339696. doi: 10.3389/fmicb.2024.1339696. eCollection 2024. Front Microbiol. 2024. PMID: 38328431 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

