Cell-free transcription-translation system: a dual read-out assay to characterize riboswitch function
- PMID: 37409574
- PMCID: PMC10450168
- DOI: 10.1093/nar/gkad574
Cell-free transcription-translation system: a dual read-out assay to characterize riboswitch function
Abstract
Cell-free protein synthesis assays have become a valuable tool to understand transcriptional and translational processes. Here, we established a fluorescence-based coupled in vitro transcription-translation assay as a read-out system to simultaneously quantify mRNA and protein levels. We utilized the well-established quantification of the expression of shifted green fluorescent protein (sGFP) as a read-out of protein levels. In addition, we determined mRNA quantities using a fluorogenic Mango-(IV) RNA aptamer that becomes fluorescent upon binding to the fluorophore thiazole orange (TO). We utilized a Mango-(IV) RNA aptamer system comprising four subsequent Mango-(IV) RNA aptamer elements with improved sensitivity by building Mango arrays. The design of this reporter assay resulted in a sensitive read-out with a high signal-to-noise ratio, allowing us to monitor transcription and translation time courses in cell-free assays with continuous monitoring of fluorescence changes as well as snapshots of the reaction. Furthermore, we applied this dual read-out assay to investigate the function of thiamine-sensing riboswitches thiM and thiC from Escherichia coli and the adenine-sensing riboswitch ASW from Vibrio vulnificus and pbuE from Bacillus subtilis, which represent transcriptional and translational on- and off-riboswitches, respectively. This approach enabled a microplate-based application, a valuable addition to the toolbox for high-throughput screening of riboswitch function.
© The Author(s) 2023. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


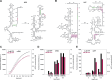

References
-
- Matthaei H., Nirenberg M.W. The dependence of cell-free protein synthesis in E. coli upon RNA prepared from ribosomes. Biochem. Biophys. Res. Commun. 1961; 4:404–408. - PubMed
-
- Katzen F., Chang G., Kudlicki W. The past, present and future of cell-free protein synthesis. Trends Biotechnol. 2005; 23:150–156. - PubMed
-
- He M. Cell-free protein synthesis: applications in proteomics and biotechnology. N. Biotechnol. 2008; 25:126–132. - PubMed
-
- Spirin A.S. High-throughput cell-free systems for synthesis of functionally active proteins. Trends Biotechnol. 2004; 22:538–545. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

