Commensal bacteria weaken the intestinal barrier by suppressing epithelial neuropilin-1 and Hedgehog signaling
- PMID: 37414930
- PMCID: PMC10365997
- DOI: 10.1038/s42255-023-00828-5
Commensal bacteria weaken the intestinal barrier by suppressing epithelial neuropilin-1 and Hedgehog signaling
Erratum in
-
Author Correction: Commensal bacteria weaken the intestinal barrier by suppressing epithelial neuropilin-1 and Hedgehog signaling.Nat Metab. 2023 Sep;5(9):1639. doi: 10.1038/s42255-023-00901-z. Nat Metab. 2023. PMID: 37697057 Free PMC article. No abstract available.
Abstract
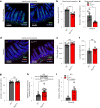
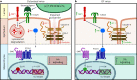
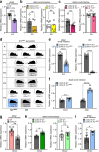
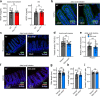
The gut microbiota influences intestinal barrier integrity through mechanisms that are incompletely understood. Here we show that the commensal microbiota weakens the intestinal barrier by suppressing epithelial neuropilin-1 (NRP1) and Hedgehog (Hh) signaling. Microbial colonization of germ-free mice dampens signaling of the intestinal Hh pathway through epithelial Toll-like receptor (TLR)-2, resulting in decreased epithelial NRP1 protein levels. Following activation via TLR2/TLR6, epithelial NRP1, a positive-feedback regulator of Hh signaling, is lysosomally degraded. Conversely, elevated epithelial NRP1 levels in germ-free mice are associated with a strengthened gut barrier. Functionally, intestinal epithelial cell-specific Nrp1 deficiency (Nrp1ΔIEC) results in decreased Hh pathway activity and a weakened gut barrier. In addition, Nrp1ΔIEC mice have a reduced density of capillary networks in their small intestinal villus structures. Collectively, our results reveal a role for the commensal microbiota and epithelial NRP1 signaling in the regulation of intestinal barrier function through postnatal control of Hh signaling.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Abrams GD, Bauer H, Sprinz H. Influence of the normal flora on mucosal morphology and cellular renewal in the ileum. A comparison of germ-free and conventional mice. Lab. Invest. 1963;12:355–364. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

