Marked variations in gut microbial diversity, functions, and disease risk between wild and captive alpine musk deer
- PMID: 37421471
- PMCID: PMC10390370
- DOI: 10.1007/s00253-023-12675-1
Marked variations in gut microbial diversity, functions, and disease risk between wild and captive alpine musk deer
Abstract
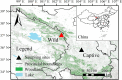
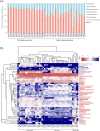
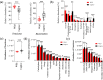
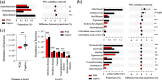
Maintaining a healthy status is crucial for the successful captive breeding of endangered alpine musk deer (Moschus chrysogaster, AMD), and captive breeding programs are beneficial to the ex-situ conservation and wild population recovery of this species. Meanwhile, the gut microbiota is essential for host health, survival, and environmental adaptation. However, changes in feeding environment and food can affect the composition and function of gut microbiota in musk deer, ultimately impacting their health and adaptation. Therefore, regulating the health status of wild and captive AMD through a non-invasive method that targets gut microbiota is a promising approach. Here, 16S rRNA gene sequencing was employed to reveal the composition and functional variations between wild (N = 23) and captive (N = 25) AMD populations. The results indicated that the gut microbiota of wild AMD exhibited significantly higher alpha diversity (P < 0.001) and greater abundance of the phylum Firmicutes, as well as several dominant genera, including UCG-005, Christensenellaceae R7 group, Monoglobus, Ruminococcus, and Roseburia (P < 0.05), compared to captive AMD. These findings suggest that the wild AMD may possess more effective nutrient absorption and utilization, a more stable intestinal microecology, and better adaption to the complex natural environment. The captive individuals displayed higher metabolic functions with an increased abundance of the phylum Bacteroidetes and certain dominant genera, including Bacteroides, Rikenellaceae RC9 gut group, NK4A214 group, and Alistipes (P < 0.05), which contributed to the metabolic activities of various nutrients. Furthermore, captive AMD showed a higher level of 11 potential opportunistic pathogens and a greater enrichment of disease-related functions compared to wild AMD, indicating that wild musk deer have a lower risk of intestinal diseases and more stable intestinal structure in comparison to captive populations. These findings can serve as a valuable theoretical foundation for promoting the healthy breeding of musk deer and as a guide for evaluating the health of wild-released and reintroduced musk deer in the future. KEY POINTS: • Wild and captive AMD exhibit contrasting gut microbial diversity and certain functions. • With higher diversity, certain bacteria aid wild AMD's adaptation to complex habitats. • Higher potential pathogens and functions increase disease risk in captive AMD.
Keywords: 16S rRNA gene sequencing; Alpine musk deer; Core microbiome; Gut microbial function; Opportunistic pathogens.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


References
MeSH terms
Substances
Grants and funding
- 2023-ZJ-952Q/Natural Science Foundation of Qinghai
- 2023-ZJ-901T/Natural Science Foundation of Qinghai
- U20A2012/National Natural Science Foundation of China
- XDA23060602/Strategic Priority Research Program of the Chinese Academy of Sciences
- LHZX-2020-01/Joint Grant from Chinese Academy of Sciences-People's Government of Qinghai Province on Sanjiangyuan National Park
LinkOut - more resources
Full Text Sources

