This is a preprint.
Identification of small molecule inhibitors of G3BP-driven stress granule formation
- PMID: 37425931
- PMCID: PMC10327151
- DOI: 10.1101/2023.06.27.546770
Identification of small molecule inhibitors of G3BP-driven stress granule formation
Update in
-
Identification of small molecule inhibitors of G3BP-driven stress granule formation.J Cell Biol. 2024 Mar 4;223(3):e202308083. doi: 10.1083/jcb.202308083. Epub 2024 Jan 29. J Cell Biol. 2024. PMID: 38284934 Free PMC article.
Abstract
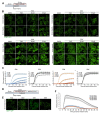
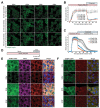
Stress granule formation is triggered by the release of mRNAs from polysomes and is promoted by the action of the paralogs G3BP1 and G3BP2. G3BP1/2 proteins bind mRNAs and thereby promote the condensation of mRNPs into stress granules. Stress granules have been implicated in several disease states, including cancer and neurodegeneration. Consequently, compounds that limit stress granule formation or promote their dissolution have potential as both experimental tools and novel therapeutics. Herein, we describe two small molecules, referred to as G3BP inhibitor a and b (G3Ia and G3Ib), designed to bind to a specific pocket in G3BP1/2 that is known to be targeted by viral inhibitors of G3BP1/2 function. In addition to disrupting co-condensation of RNA, G3BP1, and caprin 1 in vitro, these compounds inhibit stress granule formation in cells treated prior to or concurrent with stress, and dissolve pre-existing stress granules when added to cells after stress granule formation. These effects are consistent across multiple cell types and a variety of initiating stressors. Thus, these compounds represent ideal tools to probe the biology of stress granules and hold promise for therapeutic interventions designed to modulate stress granule formation.
Conflict of interest statement
Conflict of interest: JPT is a consultant for Nido Biosciences.
Figures



References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
