Accelerating bioinformatics implementation in public health
- PMID: 37428142
- PMCID: PMC10438813
- DOI: 10.1099/mgen.0.001051
Accelerating bioinformatics implementation in public health
Abstract
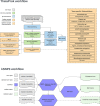
We have adopted an open bioinformatics ecosystem to address the challenges of bioinformatics implementation in public health laboratories (PHLs). Bioinformatics implementation for public health requires practitioners to undertake standardized bioinformatic analyses and generate reproducible, validated and auditable results. It is essential that data storage and analysis are scalable, portable and secure, and that implementation of bioinformatics fits within the operational constraints of the laboratory. We address these requirements using Terra, a web-based data analysis platform with a graphical user interface connecting users to bioinformatics analyses without the use of code. We have developed bioinformatics workflows for use with Terra that specifically meet the needs of public health practitioners. These Theiagen workflows perform genome assembly, quality control, and characterization, as well as construction of phylogeny for insights into genomic epidemiology. Additonally, these workflows use open-source containerized software and the WDL workflow language to ensure standardization and interoperability with other bioinformatics solutions, whilst being adaptable by the user. They are all open source and publicly available in Dockstore with the version-controlled code available in public GitHub repositories. They have been written to generate outputs in standardized file formats to allow for further downstream analysis and visualization with separate genomic epidemiology software. Testament to this solution meeting the requirements for bioinformatic implementation in public health, Theiagen workflows have collectively been used for over 5 million sample analyses in the last 2 years by over 90 public health laboratories in at least 40 different countries. Continued adoption of technological innovations and development of further workflows will ensure that this ecosystem continues to benefit PHLs.
Keywords: Terra; bioinformatics; epidemiology; genome; public health; sequencing.
Conflict of interest statement
Many authors are employed by Theiagen Genomics, a for-profit private company. J.R.S., K.G.L. and G.L. are owners of Theiagen Global, a for-profit private company with a reseller agreement with Seqera Labs, the developer of Nextflow Tower.
Figures


References
-
- Armstrong GL, MacCannell DR, Taylor J, Carleton HA, Neuhaus EB, et al. Pathogen Genomics in Public Health. Obstet Gynecol Surv. 2020;75:275–276. doi: 10.1097/01.ogx.0000666232.13540.20. - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

