CMOT: Cross-Modality Optimal Transport for multimodal inference
- PMID: 37434182
- PMCID: PMC10334579
- DOI: 10.1186/s13059-023-02989-8
CMOT: Cross-Modality Optimal Transport for multimodal inference
Abstract
Multimodal measurements of single-cell sequencing technologies facilitate a comprehensive understanding of specific cellular and molecular mechanisms. However, simultaneous profiling of multiple modalities of single cells is challenging, and data integration remains elusive due to missing modalities and cell-cell correspondences. To address this, we developed a computational approach, Cross-Modality Optimal Transport (CMOT), which aligns cells within available multi-modal data (source) onto a common latent space and infers missing modalities for cells from another modality (target) of mapped source cells. CMOT outperforms existing methods in various applications from developing brain, cancers to immunology, and provides biological interpretations improving cell-type or cancer classifications.
Keywords: Cross-modal inference; Multimodal data alignment; Optimal transport; Probabilistic coupling; Single-cell multi-modality; Weighted nearest neighbor.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



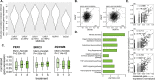

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

