Interplay between chromosomal architecture and termination of DNA replication in bacteria
- PMID: 37434703
- PMCID: PMC10331603
- DOI: 10.3389/fmicb.2023.1180848
Interplay between chromosomal architecture and termination of DNA replication in bacteria
Abstract
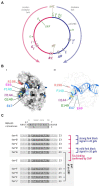
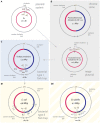
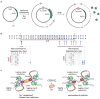
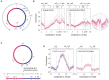
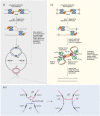
Faithful transmission of the genome from one generation to the next is key to life in all cellular organisms. In the majority of bacteria, the genome is comprised of a single circular chromosome that is normally replicated from a single origin, though additional genetic information may be encoded within much smaller extrachromosomal elements called plasmids. By contrast, the genome of a eukaryote is distributed across multiple linear chromosomes, each of which is replicated from multiple origins. The genomes of archaeal species are circular, but are predominantly replicated from multiple origins. In all three cases, replication is bidirectional and terminates when converging replication fork complexes merge and 'fuse' as replication of the chromosomal DNA is completed. While the mechanics of replication initiation are quite well understood, exactly what happens during termination is far from clear, although studies in bacterial and eukaryotic models over recent years have started to provide some insight. Bacterial models with a circular chromosome and a single bidirectional origin offer the distinct advantage that there is normally just one fusion event between two replication fork complexes as synthesis terminates. Moreover, whereas termination of replication appears to happen in many bacteria wherever forks happen to meet, termination in some bacterial species, including the well-studied bacteria Escherichia coli and Bacillus subtilis, is more restrictive and confined to a 'replication fork trap' region, making termination even more tractable. This region is defined by multiple genomic terminator (ter) sites, which, if bound by specific terminator proteins, form unidirectional fork barriers. In this review we discuss a range of experimental results highlighting how the fork fusion process can trigger significant pathologies that interfere with the successful conclusion of DNA replication, how these pathologies might be resolved in bacteria without a fork trap system and how the acquisition of a fork trap might have provided an alternative and cleaner solution, thus explaining why in bacterial species that have acquired a fork trap system, this system is remarkably well maintained. Finally, we consider how eukaryotic cells can cope with a much-increased number of termination events.
Keywords: DNA replication; DNA segregation; RecG helicase; Tus-ter complexes; bacterial chromosome dynamics; chromosomal architecture; termination of DNA replication.
Copyright © 2023 Goodall, Warecka, Hawkins and Rudolph.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Termination of DNA replication at Tus-ter barriers results in under-replication of template DNA.J Biol Chem. 2021 Dec;297(6):101409. doi: 10.1016/j.jbc.2021.101409. Epub 2021 Nov 12. J Biol Chem. 2021. PMID: 34780717 Free PMC article.
-
A Fork Trap in the Chromosomal Termination Area Is Highly Conserved across All Escherichia coli Phylogenetic Groups.Int J Mol Sci. 2021 Jul 25;22(15):7928. doi: 10.3390/ijms22157928. Int J Mol Sci. 2021. PMID: 34360694 Free PMC article.
-
Chromosomal over-replication in Escherichia coli recG cells is triggered by replication fork fusion and amplified if replichore symmetry is disturbed.Nucleic Acids Res. 2018 Sep 6;46(15):7701-7715. doi: 10.1093/nar/gky566. Nucleic Acids Res. 2018. PMID: 29982635 Free PMC article.
-
Replication Termination: Containing Fork Fusion-Mediated Pathologies in Escherichia coli.Genes (Basel). 2016 Jul 25;7(8):40. doi: 10.3390/genes7080040. Genes (Basel). 2016. PMID: 27463728 Free PMC article. Review.
-
Too Much of a Good Thing: How Ectopic DNA Replication Affects Bacterial Replication Dynamics.Front Microbiol. 2020 Apr 15;11:534. doi: 10.3389/fmicb.2020.00534. eCollection 2020. Front Microbiol. 2020. PMID: 32351461 Free PMC article. Review.
Cited by
-
Spatio-temporal organization of the E. coli chromosome from base to cellular length scales.EcoSal Plus. 2024 Dec 12;12(1):eesp00012022. doi: 10.1128/ecosalplus.esp-0001-2022. Epub 2024 Jun 12. EcoSal Plus. 2024. PMID: 38864557 Free PMC article. Review.
-
Escherichia coli DNA replication: the old model organism still holds many surprises.FEMS Microbiol Rev. 2024 Jun 20;48(4):fuae018. doi: 10.1093/femsre/fuae018. FEMS Microbiol Rev. 2024. PMID: 38982189 Free PMC article. Review.
References
Publication types
LinkOut - more resources
Full Text Sources

