Proximity-dependent recruitment of Polycomb repressive complexes by the lncRNA Airn
- PMID: 37436897
- PMCID: PMC10441531
- DOI: 10.1016/j.celrep.2023.112803
Proximity-dependent recruitment of Polycomb repressive complexes by the lncRNA Airn
Abstract
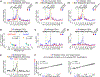
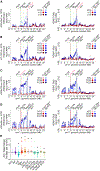
During mouse embryogenesis, expression of the long non-coding RNA (lncRNA) Airn leads to gene repression and recruitment of Polycomb repressive complexes (PRCs) to varying extents over a 15-Mb domain. The mechanisms remain unclear. Using high-resolution approaches, we show in mouse trophoblast stem cells that Airn expression induces long-range changes to chromatin architecture that coincide with PRC-directed modifications and center around CpG island promoters that contact the Airn locus even in the absence of Airn expression. Intensity of contact between the Airn lncRNA and chromatin correlated with underlying intensity of PRC recruitment and PRC-directed modifications. Deletion of CpG islands that contact the Airn locus altered long-distance repression and PRC activity in a manner that correlated with changes in chromatin architecture. Our data imply that the extent to which Airn expression recruits PRCs to chromatin is controlled by DNA regulatory elements that modulate proximity of the Airn lncRNA product to its target DNA.
Keywords: Airn; CHART-seq; CP: Molecular biology; ChIP-seq; Hi-C; Xist; chromatin archtecture; epigenetics; long noncoding RNA; polycomb repressive complex; transcription.
Copyright © 2023 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures





References
-
- Huang R, Jaritz M, Guenzl P, Vlatkovic I, Sommer A, Tamir IM, Marks H, Klampfl T, Kralovics R, Stunnenberg HG, et al. (2011). An RNA-Seq strategy to detect the complete coding and non-coding transcriptome including full-length imprinted macro ncRNAs. PLoS One 6, e27288. 10.1371/journal.pone.0027288. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

