Predicting Structural Susceptibility of Proteins to Proteolytic Processing
- PMID: 37445939
- PMCID: PMC10342023
- DOI: 10.3390/ijms241310761
Predicting Structural Susceptibility of Proteins to Proteolytic Processing
Abstract
The importance of 3D protein structure in proteolytic processing is well known. However, despite the plethora of existing methods for predicting proteolytic sites, only a few of them utilize the structural features of potential substrates as predictors. Moreover, to our knowledge, there is currently no method available for predicting the structural susceptibility of protein regions to proteolysis. We developed such a method using data from CutDB, a database that contains experimentally verified proteolytic events. For prediction, we utilized structural features that have been shown to influence proteolysis in earlier studies, such as solvent accessibility, secondary structure, and temperature factor. Additionally, we introduced new structural features, including length of protruded loops and flexibility of protein termini. To maximize the prediction quality of the method, we carefully curated the training set, selected an appropriate machine learning method, and sampled negative examples to determine the optimal positive-to-negative class size ratio. We demonstrated that combining our method with models of protease primary specificity can outperform existing bioinformatics methods for the prediction of proteolytic sites. We also discussed the possibility of utilizing this method for bioinformatics prediction of other post-translational modifications.
Keywords: protease substrates; proteases; regulatory proteolysis; substrate identification.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
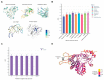
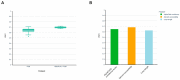

Similar articles
-
Sequence-derived structural features driving proteolytic processing.Proteomics. 2014 Jan;14(1):42-50. doi: 10.1002/pmic.201300416. Epub 2013 Dec 11. Proteomics. 2014. PMID: 24227478 Free PMC article.
-
Structural determinants of limited proteolysis.J Proteome Res. 2011 Aug 5;10(8):3642-51. doi: 10.1021/pr200271w. Epub 2011 Jul 8. J Proteome Res. 2011. PMID: 21682278 Free PMC article.
-
PROSPER: an integrated feature-based tool for predicting protease substrate cleavage sites.PLoS One. 2012;7(11):e50300. doi: 10.1371/journal.pone.0050300. Epub 2012 Nov 29. PLoS One. 2012. PMID: 23209700 Free PMC article.
-
iProt-Sub: a comprehensive package for accurately mapping and predicting protease-specific substrates and cleavage sites.Brief Bioinform. 2019 Mar 25;20(2):638-658. doi: 10.1093/bib/bby028. Brief Bioinform. 2019. PMID: 29897410 Free PMC article. Review.
-
Protein TAILS: when termini tell tales of proteolysis and function.Curr Opin Chem Biol. 2013 Feb;17(1):73-82. doi: 10.1016/j.cbpa.2012.11.025. Epub 2013 Jan 6. Curr Opin Chem Biol. 2013. PMID: 23298954 Review.
Cited by
-
Genome-wide bioinformatics analysis of human protease capacity for proteolytic cleavage of the SARS-CoV-2 spike glycoprotein.Microbiol Spectr. 2024 Feb 6;12(2):e0353023. doi: 10.1128/spectrum.03530-23. Epub 2024 Jan 8. Microbiol Spectr. 2024. PMID: 38189333 Free PMC article.
-
Identification of pancreatic cancer-specific protease substrates for protease-dependent targeted delivery.Oncogenesis. 2024 Nov 20;13(1):40. doi: 10.1038/s41389-024-00542-1. Oncogenesis. 2024. PMID: 39567504 Free PMC article.
References
-
- Ratnikov B.I., Cieplak P., Gramatikoff K., Pierce J., Eroshkin A., Igarashi Y., Kazanov M., Sun Q., Godzik A., Osterman A., et al. Basis for Substrate Recognition and Distinction by Matrix Metalloproteinases. Proc. Natl. Acad. Sci. USA. 2014;111:E4148–E4155. doi: 10.1073/pnas.1406134111. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

