Vps4a Regulates Autophagic Flux to Prevent Hypertrophic Cardiomyopathy
- PMID: 37445978
- PMCID: PMC10341959
- DOI: 10.3390/ijms241310800
Vps4a Regulates Autophagic Flux to Prevent Hypertrophic Cardiomyopathy
Abstract
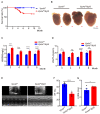
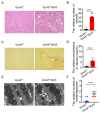
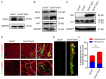
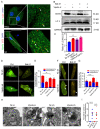
Autophagy has stabilizing functions for cardiomyocytes. Recent studies indicate that an impairment in the autophagy pathway can seriously affect morphology and function, potentially leading to heart failure. However, the role and the underlying mechanism of the endosomal sorting complex required for transport (ESCRT) family protein, in particular the AAA-ATPase vacuolar protein sorting 4a (Vps4a), in regulating myocardial autophagy remains unclear. In the present study, cardiomyocyte-specific Vps4a knockout mice were generated by crossing Vps4aflox/flox (Vps4afl/fl) with Myh6-cre transgenic mice. As a result, we observed a partially dilated left ventricular (LV) chamber, a significant increase in heart weight to body weight ratio (HW/BW), and heart weight to tibial length ratio (HW/TL), hypertrophic cardiomyopathy and early lethality starting at 3 months of age. Hematoxylin-eosin (HE), immunofluorescence assay (IFA), and Western blot (WB) revealed autophagosome accumulation in cardiomyocytes. A transcriptome-based analysis and autophagic flux tracking by AAV-RFP-GFP-LC3 showed that the autophagic flux was blocked in Vps4a knockout cardiomyocytes. In addition, we provided in vitro evidence demonstrating that Vps4a and LC3 were partially co-localized in cardiomyocytes, and the knockdown of Vps4a led to the accumulation of autophagosomes in cardiomyocytes. Similarly, the transfection of cardiomyocytes with adenovirus (Adv) mCherry-GFP-LC3 further indicated that the autophagic flux was blocked in cells with deficient levels of Vps4a. Finally, an electron microscope (EM) showed that the compromised sealing of autophagosome blocked the autophagic flux in Vps4a-depleted cardiomyocytes. These findings revealed that Vps4a contributed to the sealing of autophagosomes in cardiomyocytes. Therefore, we demonstrated that Vps4a deletion could block the autophagic flux, leading to the accumulation of degradation substances and compromised cardiac function. Overall, this study provides insights into a new theoretical basis for which autophagy may represent a therapeutic target for cardiovascular diseases.
Keywords: ESCRT; Vps4a; autophagy; heart; heart failure.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

Similar articles
-
Downregulation of LAPTM4B Contributes to the Impairment of the Autophagic Flux via Unopposed Activation of mTORC1 Signaling During Myocardial Ischemia/Reperfusion Injury.Circ Res. 2020 Sep 11;127(7):e148-e165. doi: 10.1161/CIRCRESAHA.119.316388. Epub 2020 Jul 22. Circ Res. 2020. PMID: 32693673
-
Vps4a Mediates a Unified Membrane Repair Machinery to Attenuate Ischemia/Reperfusion Injury.Circ Res. 2025 Jan 31;136(3):279-296. doi: 10.1161/CIRCRESAHA.124.325290. Epub 2025 Jan 7. Circ Res. 2025. PMID: 39764631
-
Overexpression of FNTB and the activation of Ras induce hypertrophy and promote apoptosis and autophagic cell death in cardiomyocytes.J Cell Mol Med. 2020 Aug;24(16):8998-9011. doi: 10.1111/jcmm.15533. Epub 2020 Jun 24. J Cell Mol Med. 2020. PMID: 32579303 Free PMC article.
-
Sustained over-expression of calpain-2 induces age-dependent dilated cardiomyopathy in mice through aberrant autophagy.Acta Pharmacol Sin. 2022 Nov;43(11):2873-2884. doi: 10.1038/s41401-022-00965-9. Epub 2022 Aug 19. Acta Pharmacol Sin. 2022. PMID: 35986214 Free PMC article.
-
The Molecular Mechanism and Therapeutic Application of Autophagy for Urological Disease.Int J Mol Sci. 2023 Oct 4;24(19):14887. doi: 10.3390/ijms241914887. Int J Mol Sci. 2023. PMID: 37834333 Free PMC article. Review.
Cited by
-
EPHB2 Promotes the Progression of Oral Squamous Cell Carcinoma Cells Through the Activation of VPS4A-Mediated Autophagy.Cancer Sci. 2025 May;116(5):1308-1323. doi: 10.1111/cas.70033. Epub 2025 Feb 27. Cancer Sci. 2025. PMID: 40017157 Free PMC article.
-
Identification and validation of autophagy‑related genes in hypertrophic cardiomyopathy.Exp Ther Med. 2024 Sep 25;28(6):440. doi: 10.3892/etm.2024.12729. eCollection 2024 Dec. Exp Ther Med. 2024. PMID: 39355520 Free PMC article.
-
Urine Proteomic Signatures of Mild Hypothermia Treatment in Cerebral Ischemia-Reperfusion Injury in Rats.Cell Mol Neurobiol. 2024 Jun 5;44(1):49. doi: 10.1007/s10571-024-01483-4. Cell Mol Neurobiol. 2024. PMID: 38836960 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

