Wood fibers are a crucial microhabitat for cellulose- and xylan- degrading bacteria in the hindgut of the wood-feeding beetle Odontotaenius disjunctus
- PMID: 37448580
- PMCID: PMC10338082
- DOI: 10.3389/fmicb.2023.1173696
Wood fibers are a crucial microhabitat for cellulose- and xylan- degrading bacteria in the hindgut of the wood-feeding beetle Odontotaenius disjunctus
Abstract
Introduction: Wood digestion in insects relies on the maintenance of a mosaic of numerous microhabitats, each colonized by distinct microbiomes. Understanding the division of digestive labor between these microhabitats- is central to understanding the physiology and evolution of symbiotic wood digestion. A microhabitat that has emerged to be of direct relevance to the process of lignocellulose digestion is the surface of ingested plant material. Wood particles in the guts of some termites are colonized by a specialized bacterial fiber-digesting microbiome, but whether this represents a widespread strategy among insect lineages that have independently evolved wood-feeding remains an open question.
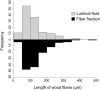
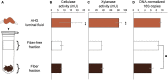
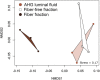
Methods: In this study, we investigated the bacterial communities specifically associated with wood fibers in the gut of the passalid beetle Odontotaenius disjunctus. We developed a Percoll-based centrifugation method to isolate and enrich the wood particles from the anterior hindgut, allowing us to access the wood fibers and their associated microbiome. We then performed assays of enzyme activity and used short-read and long-read amplicon sequencing of the 16S rRNA gene to identify the composition of the fiber-associated microbiome.
Results: Our assays demonstrated that the anterior hindgut, which houses a majority of the bacterial load, is an important site for lignocellulose digestion. Wood particles enriched from the anterior hindgut contribute to a large proportion of the total enzyme activity. The sequencing revealed that O. disjunctus, like termites, harbors a distinct fiber-associated microbiome, but notably, its community is enriched in insect-specific groups of Lactococcus and Turicibacter.
Discussion: Our study underscores the importance of microhabitats in fostering the complex symbiotic relationships between wood-feeding insects and their microbiomes. The discovery of distinct fiber-digesting symbionts in O. disjunctus, compared to termites, highlights the diverse evolutionary paths insects have taken to adapt to a challenging diet.
Keywords: Passalidae; beetles; gut microbiomes; lignocellulose; symbiotic digestion.
Copyright © 2023 Schwarz, Beza-Beza and Mikaelyan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



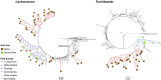
References
-
- Bauer E., Lampert N., Mikaelyan A., Köhler T., Maekawa K., Brune A. (2015). Physicochemical conditions, metabolites and community structure of the bacterial microbiota in the gut of wood-feeding cockroaches (Blaberidae: Panesthiinae). FEMS Microbiol. Ecol. 91 1–14. 10.1093/femsec/fiu028 - DOI - PubMed
-
- Brulc J. M., Antonopoulos D. A., Berg Miller M. E., Wilson M. K., Yannarell A. C., Dinsdale E. A., et al. (2009). Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc. Natl. Acad. Sci. U.S.A. 106 1948–1953. 10.1073/pnas.0806191105 - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

