Lactate-dependent transcriptional regulation controls mammalian eye morphogenesis
- PMID: 37452018
- PMCID: PMC10349100
- DOI: 10.1038/s41467-023-39672-2
Lactate-dependent transcriptional regulation controls mammalian eye morphogenesis
Abstract
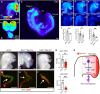
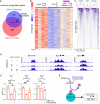
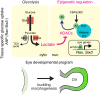
Mammalian retinal metabolism favors aerobic glycolysis. However, the role of glycolytic metabolism in retinal morphogenesis remains unknown. We report that aerobic glycolysis is necessary for the early stages of retinal development. Taking advantage of an unbiased approach that combines the use of eye organoids and single-cell RNA sequencing, we identify specific glucose transporters and glycolytic genes in retinal progenitors. Next, we determine that the optic vesicle territory of mouse embryos displays elevated levels of glycolytic activity. At the functional level, we show that removal of Glucose transporter 1 and Lactate dehydrogenase A gene activity from developing retinal progenitors arrests eye morphogenesis. Surprisingly, we uncover that lactate-mediated upregulation of key eye-field transcription factors is controlled by the epigenetic modification of histone H3 acetylation through histone deacetylase activity. Our results identify an unexpected bioenergetic independent role of lactate as a signaling molecule necessary for mammalian eye morphogenesis.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

