Transcriptional insights into Chlorella sp. ABC-001: a comparative study of carbon fixation and lipid synthesis under different CO2 conditions
- PMID: 37454088
- PMCID: PMC10350272
- DOI: 10.1186/s13068-023-02358-4
Transcriptional insights into Chlorella sp. ABC-001: a comparative study of carbon fixation and lipid synthesis under different CO2 conditions
Abstract
Background: Microalgae's low tolerance to high CO2 concentrations presents a significant challenge for its industrial application, especially when considering the utilization of industrial exhaust gas streams with high CO2 content-an economically and environmentally attractive option. Therefore, the objectives of this study were to investigate the metabolic changes in carbon fixation and lipid accumulation of microalgae under ambient air and high CO2 conditions, deepen our understanding of the molecular mechanisms driving these processes, and identify potential target genes for metabolic engineering in microalgae. To accomplish these goals, we conducted a transcriptomic analysis of the high CO2-tolerant strain, Chlorella sp. ABC-001, under two different carbon dioxide levels (ambient air and 10% CO2) and at various growth phases.
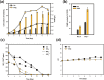
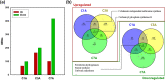
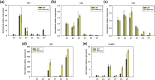
Results: Cells cultivated with 10% CO2 exhibited significantly better growth and lipid accumulation rates, achieving up to 2.5-fold higher cell density and twice the lipid content by day 7. To understand the relationship between CO2 concentrations and phenotypes, transcriptomic analysis was conducted across different CO2 conditions and growth phases. According to the analysis of differentially expressed genes and gene ontology, Chlorella sp. ABC-001 exhibited the development of chloroplast organelles during the early exponential phase under high CO2 conditions, resulting in improved CO2 fixation and enhanced photosynthesis. Cobalamin-independent methionine synthase expression was also significantly elevated during the early growth stage, likely contributing to the methionine supply required for various metabolic activities and active proliferation. Conversely, the cells showed sustained repression of carbonic anhydrase and ferredoxin hydrogenase, involved in the carbon concentrating mechanism, throughout the cultivation period under high CO2 conditions. This study also delved into the transcriptomic profiles in the Calvin cycle, nitrogen reductase, and lipid synthesis. Particularly, Chlorella sp. ABC-001 showed high expression levels of genes involved in lipid synthesis, such as glycerol-3-phosphate dehydrogenase and phospholipid-diacylglycerol acyltransferase. These findings suggest potential targets for metabolic engineering aimed at enhancing lipid production in microalgae.
Conclusions: We expect that our findings will help understand the carbon concentrating mechanism, photosynthesis, nitrogen assimilation, and lipid accumulation metabolisms of green algae according to CO2 concentrations. This study also provides insights into systems metabolic engineering of microalgae for improved performance in the future.
Keywords: CO2 fixation; Chlorella; Lipid accumulation; Microalgae; Transcriptomic analysis.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



Similar articles
-
The regulating mechanisms of CO2 fixation and carbon allocations of two Chlorella sp. strains in response to high CO2 levels.Chemosphere. 2020 May;247:125814. doi: 10.1016/j.chemosphere.2020.125814. Epub 2020 Jan 7. Chemosphere. 2020. PMID: 31927186
-
A biorefinery for valorization of industrial waste-water and flue gas by microalgae for waste mitigation, carbon-dioxide sequestration and algal biomass production.Sci Total Environ. 2019 Oct 20;688:129-135. doi: 10.1016/j.scitotenv.2019.06.024. Epub 2019 Jun 6. Sci Total Environ. 2019. PMID: 31229810
-
Enhanced lipid accumulation of photoautotrophic microalgae by high-dose CO2 mimics a heterotrophic characterization.World J Microbiol Biotechnol. 2016 Jan;32(1):9. doi: 10.1007/s11274-015-1963-6. Epub 2015 Dec 28. World J Microbiol Biotechnol. 2016. PMID: 26712624
-
Monitoring lipids profile, CO2 fixation, and water recyclability for the economic viability of microalgae Chlorella vulgaris cultivation at different initial nitrogen.J Biotechnol. 2022 Feb 10;345:30-39. doi: 10.1016/j.jbiotec.2021.12.014. Epub 2022 Jan 4. J Biotechnol. 2022. PMID: 34995559 Review.
-
Molecular characterization of CO2 sequestration and assimilation in microalgae and its biotechnological applications.Bioresour Technol. 2017 Nov;244(Pt 2):1207-1215. doi: 10.1016/j.biortech.2017.05.199. Epub 2017 Jun 1. Bioresour Technol. 2017. PMID: 28606753 Review.
Cited by
-
Usage of Chlorella and diverse microalgae for CO2 capture - towards a bioenergy revolution.Front Bioeng Biotechnol. 2024 Aug 20;12:1387519. doi: 10.3389/fbioe.2024.1387519. eCollection 2024. Front Bioeng Biotechnol. 2024. PMID: 39229458 Free PMC article. Review.
-
Induced Resistance against Pseudomonas syringae pv. lachrymans in Cucumber by Spraying Cell-Free Microalgae Supernatant.Plant Pathol J. 2025 Jun;41(3):321-329. doi: 10.5423/PPJ.FT.02.2025.0028. Epub 2025 Jun 1. Plant Pathol J. 2025. PMID: 40468879 Free PMC article.
References
-
- Singh UB, Ahluwalia AS. Microalgae: a promising tool for carbon sequestration. Mitig Adapt Strat Glob Change. 2013;18(1):73–95. doi: 10.1007/s11027-012-9393-3. - DOI
-
- Packer M. Algal capture of carbon dioxide; biomass generation as a tool for greenhouse gas mitigation with reference to New Zealand energy strategy and policy. Energy Policy. 2009;37(9):3428–3437. doi: 10.1016/j.enpol.2008.12.025. - DOI
-
- International Energy Agency. State of Technology Review on Algae Bioenergy. 2017. https://www.ieabioenergy.com/wp-content/uploads/2017/01/IEA-Bioenergy-Al...
-
- Morales M, Sánchez L, Revah S. The impact of environmental factors on carbon dioxide fixation by microalgae. FEMS Microbiol Lett. 2018;365(3). - PubMed
-
- Gerotto C, Norici A, Giordano M. Toward enhanced fixation of CO2 in aquatic biomass: focus on microalgae. Front Energy Res. 2020 doi: 10.3389/fenrg.2020.00213. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
