Attenuation hotspots in neurotropic human astroviruses
- PMID: 37459343
- PMCID: PMC10374088
- DOI: 10.1371/journal.pbio.3001815
Attenuation hotspots in neurotropic human astroviruses
Abstract
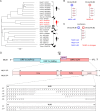
During the last decade, the detection of neurotropic astroviruses has increased dramatically. The MLB genogroup of astroviruses represents a genetically distinct group of zoonotic astroviruses associated with gastroenteritis and severe neurological complications in young children, the immunocompromised, and the elderly. Using different virus evolution approaches, we identified dispensable regions in the 3' end of the capsid-coding region responsible for attenuation of MLB astroviruses in susceptible cell lines. To create recombinant viruses with identified deletions, MLB reverse genetics (RG) and replicon systems were developed. Recombinant truncated MLB viruses resulted in imbalanced RNA synthesis and strong attenuation in iPSC-derived neuronal cultures confirming the location of neurotropism determinants. This approach can be used for the development of vaccine candidates using attenuated astroviruses that infect humans, livestock animals, and poultry.
Copyright: © 2023 Ali et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

