Panax notoginseng transcription factor WRKY15 modulates resistance to Fusarium solani by up-regulating osmotin-like protein expression and inducing JA/SA signaling pathways
- PMID: 37460949
- PMCID: PMC10351173
- DOI: 10.1186/s12870-023-04373-x
Panax notoginseng transcription factor WRKY15 modulates resistance to Fusarium solani by up-regulating osmotin-like protein expression and inducing JA/SA signaling pathways
Abstract
Background: Panax notoginseng (Burk) F. H. Chen is a valuable traditional Chinese medicinal plant, but its commercial production is seriously affected by root rot caused by some pathogenic fungi, including Fusarium solani. Nevertheless, the genetic breeding for disease resistance of P. notoginseng remains limited. The WRKY transcription factors have been revealed to play important roles in plant defense responses, which might provide an inspiration for resistance improvement in P. notoginseng.
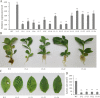
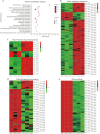
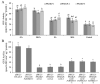
Results: In this study, the regulatory mechanism of transcription factor PnWRKY15 on P. notoginseng resistance to F. solani infection was revealed. The suppressed expression of PnWRKY15 via RNA interference increased the sensitivity of P. notoginseng to F. solani and decreased the expression levels of some defense-related genes, including PnOLP1, which encodes an osmotin-like protein that confers resistance to F. solani. Ectopic expression of PnWRKY15 in the model plant tobacco significantly enhanced the resistance to F. solani. Moreover, the transcriptome sequencing analysis discovered that some pathogenesis-related genes were expressed at higher levels in the PnWRKY15-overexpressing tobacco than that in the wild-type tobacco. In addition, the jasmonic acid (JA) and salicylic acid (SA) signaling pathways were evidently induced by PnWRKY15-overexpression, that was evidenced by that the JA and SA contents were significantly higher in the PnWRKY15-overexpressing tobacco than that in the wild-type. Furthermore, PnWRKY15, which was localized in the nucleus, can trans-activate and up-regulate PnOLP1 expression according to the EMSA, yeast one-hybrid and co-expression assays.
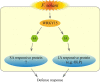
Conclusions: PnWRKY15 contributes to P. notoginseng resistance to F. solani by up-regulating the expression of resistance-related gene PnOLP1 and activating JA/SA signaling pathways. These findings will help to further elucidate the transcriptional regulatory mechanism associated with the P. notoginseng defense response to F. solani.
Keywords: Fusarium solani; Jasmonic acid; Osmotin; Panax notoginseng; Salicylic acid; WRKY transcription factor.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Osmotin-Like Protein Gene from Panax notoginseng Is Regulated by Jasmonic Acid and Involved in Defense Responses to Fusarium solani.Phytopathology. 2020 Aug;110(8):1419-1427. doi: 10.1094/PHYTO-11-19-0410-R. Epub 2020 Jun 24. Phytopathology. 2020. PMID: 32301678
-
An MYB Transcription Factor Modulates Panax notoginseng Resistance Against the Root Rot Pathogen Fusarium solani by Regulating the Jasmonate Acid Signaling Pathway and Photosynthesis.Phytopathology. 2022 Jun;112(6):1323-1334. doi: 10.1094/PHYTO-07-21-0283-R. Epub 2022 Apr 29. Phytopathology. 2022. PMID: 34844417
-
A transcriptome analysis uncovers Panax notoginseng resistance to Fusarium solani induced by methyl jasmonate.Genes Genomics. 2019 Dec;41(12):1383-1396. doi: 10.1007/s13258-019-00865-z. Epub 2019 Sep 6. Genes Genomics. 2019. PMID: 31493262
-
Molecular insights into the jasmonate signaling and associated defense responses against wilt caused by Fusarium oxysporum.Plant Physiol Biochem. 2022 Mar 1;174:22-34. doi: 10.1016/j.plaphy.2022.01.032. Epub 2022 Jan 29. Plant Physiol Biochem. 2022. PMID: 35121482 Review.
-
Pea-Fusarium solani interactions contributions of a system toward understanding disease resistance.Phytopathology. 2008 Apr;98(4):372-9. doi: 10.1094/PHYTO-98-4-0372. Phytopathology. 2008. PMID: 18944184 Review.
Cited by
-
Genome-wide identification and characterization of WRKYs family involved in responses to Cylindrocarpon destructans in Panax notoginseng.BMC Genomics. 2025 Feb 3;26(1):104. doi: 10.1186/s12864-025-11280-y. BMC Genomics. 2025. PMID: 39901065 Free PMC article.
-
Integrated multi-omics investigation revealed the importance of phenylpropanoid metabolism in the defense response of Lilium regale Wilson to fusarium wilt.Hortic Res. 2024 May 20;11(7):uhae140. doi: 10.1093/hr/uhae140. eCollection 2024 Jul. Hortic Res. 2024. PMID: 38988612 Free PMC article.
References
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

