Stability and heterogeneity in the antimicrobiota reactivity of human milk-derived immunoglobulin A
- PMID: 37462916
- PMCID: PMC10354535
- DOI: 10.1084/jem.20220839
Stability and heterogeneity in the antimicrobiota reactivity of human milk-derived immunoglobulin A
Abstract
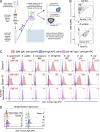
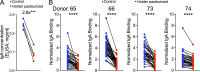
Immunoglobulin A (IgA) is secreted into breast milk and is critical for both protecting against enteric pathogens and shaping the infant intestinal microbiota. The efficacy of breast milk-derived maternal IgA (BrmIgA) is dependent upon its specificity; however, heterogeneity in BrmIgA binding ability to the infant microbiota is not known. Using a flow cytometric array, we analyzed the reactivity of BrmIgA against bacteria common to the infant microbiota and discovered substantial heterogeneity between all donors, independent of preterm or term delivery. Surprisingly, we also observed intradonor variability in the BrmIgA response to closely related bacterial isolates. Conversely, longitudinal analysis showed that the antibacterial BrmIgA reactivity was relatively stable through time, even between sequential infants, indicating that mammary gland IgA responses are durable. Together, our study demonstrates that the antibacterial BrmIgA reactivity displays interindividual heterogeneity but intraindividual stability. These findings have important implications for how breast milk shapes the development of the preterm infant microbiota and protects against necrotizing enterocolitis.
© 2023 Johnson-Hence et al.
Conflict of interest statement
Disclosures: K.P. Gopalakrishna reported a patent for Methods of screening antibodies for treating and/or preventing necrotizing enterocolitis (NEC) issued. T.W. Hand reported personal fees from Keller Postman LLC outside the submitted work; in addition, T.W. Hand had a patent to USSN: 17/341,272 pending. No other disclosures were reported.
Figures








Update of
-
Stability and heterogeneity in the anti-microbiota reactivity of human milk-derived Immunoglobulin A.bioRxiv [Preprint]. 2023 Mar 20:2023.03.16.532940. doi: 10.1101/2023.03.16.532940. bioRxiv. 2023. Update in: J Exp Med. 2023 Aug 7;220(8):e20220839. doi: 10.1084/jem.20220839. PMID: 36993366 Free PMC article. Updated. Preprint.
References
-
- Bondt, A., Dingess K.A., Hoek M., van Rijswijck D.M.H., and Heck A.J.R.. 2021. A direct MS-based approach to profile human milk secretory immunoglobulin A (IgA1) reveals donor-specific clonal repertoires with high longitudinal stability. Front. Immunol. 12:789748. 10.3389/fimmu.2021.789748 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

