Crystal structures of herbicide-detoxifying esterase reveal a lid loop affecting substrate binding and activity
- PMID: 37468532
- PMCID: PMC10356948
- DOI: 10.1038/s41467-023-40103-5
Crystal structures of herbicide-detoxifying esterase reveal a lid loop affecting substrate binding and activity
Abstract
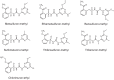
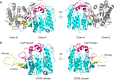
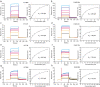
SulE, an esterase, which detoxifies a variety of sulfonylurea herbicides through de-esterification, provides an attractive approach to remove environmental sulfonylurea herbicides and develop herbicide-tolerant crops. Here, we determined the crystal structures of SulE and an activity improved mutant P44R. Structural analysis revealed that SulE is a dimer with spacious binding pocket accommodating the large sulfonylureas substrate. Particularly, SulE contains a protruding β hairpin with a lid loop covering the active site of the other subunit of the dimer. The lid loop participates in substrate recognition and binding. P44R mutation altered the lid loop flexibility, resulting in the sulfonylurea heterocyclic ring repositioning to a relative stable conformation thus leading to dramatically increased activity. Our work provides important insights into the molecular mechanism of SulE, and establish a solid foundation for further improving the enzyme activity to various sulfonylurea herbicides through rational design.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



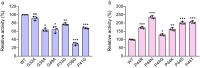
References
-
- Duggleby RG, Pang SS. Acetohydroxyacid synthase. J. Biochem. Mol. Biol. 2000;33:1–36.
-
- Saari, L. L. & Mauvais, C. J. Sulfonylurea Herbicide-resistant Crops: Herbicide-Resistant Crops (CRC Press, 2018).
-
- Blair AM, Martin TD. A review of the activity, fate and mode of action of sulfonylurea herbicides. Pestic. Sci. 1988;22:195–219. doi: 10.1002/ps.2780220303. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

