F-ATP synthase inhibitory factor 1 regulates metabolic reprogramming involving its interaction with c-Myc and PGC1α
- PMID: 37469400
- PMCID: PMC10352482
- DOI: 10.3389/fonc.2023.1207603
F-ATP synthase inhibitory factor 1 regulates metabolic reprogramming involving its interaction with c-Myc and PGC1α
Abstract
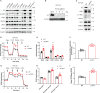
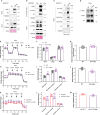
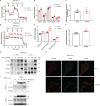
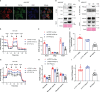
F-ATP synthase inhibitory factor 1 (IF1) is an intrinsic inhibitor of F-ATP synthase. It is known that IF1 mediates metabolic phenotypes and cell fate, yet the molecular mechanisms through which IF1 fulfills its physiological functions are not fully understood. Ablation of IF1 favors metabolic switch to oxidative metabolism from glycolysis. c-Myc and PGC1α are critical for metabolic reprogramming. This work identified that IF1 interacted with Thr-58 phosphorylated c-Myc, which might thus mediate the activity of c-Myc and promote glycolysis. The interaction of IF1 with PGC1α inhibited oxidative respiration. c-Myc and PGC1α were localized to mitochondria under mitochondrial stress in an IF1-dependent manner. Furthermore, IF1 was found to be required for the protective effect of hypoxia on c-Myc- and PGC1α-induced cell death. This study suggested that the interactions of IF1 with transcription factors c-Myc and PGC1α might be involved in IF1-regulatory metabolic reprogramming and cell fate.
Keywords: F-ATP synthase inhibitory factor 1; PGC1α; c-Myc; metabolic reprogramming; mitochondria; p-c-Myc.
Copyright © 2023 Guo and Gu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
The ATPase Inhibitory Factor 1 (IF1): A master regulator of energy metabolism and of cell survival.Biochim Biophys Acta. 2016 Aug;1857(8):1167-1182. doi: 10.1016/j.bbabio.2016.02.004. Epub 2016 Feb 12. Biochim Biophys Acta. 2016. PMID: 26876430
-
F-ATP synthase inhibitory factor 1 and mitochondria-organelle interactions: New insight and implications.Pharmacol Res. 2024 Oct;208:107393. doi: 10.1016/j.phrs.2024.107393. Epub 2024 Sep 2. Pharmacol Res. 2024. PMID: 39233058 Review.
-
A Review of the Inhibition of the Mitochondrial ATP Synthase by IF1 in vivo: Reprogramming Energy Metabolism and Inducing Mitohormesis.Front Physiol. 2018 Sep 19;9:1322. doi: 10.3389/fphys.2018.01322. eCollection 2018. Front Physiol. 2018. PMID: 30283362 Free PMC article. Review.
-
The ATPase Inhibitory Factor 1 (IF1) Contributes to the Warburg Effect and Is Regulated by Its Phosphorylation in S39 by a Protein Kinase A-like Activity.Cancers (Basel). 2024 Feb 29;16(5):1014. doi: 10.3390/cancers16051014. Cancers (Basel). 2024. PMID: 38473373 Free PMC article. Review.
-
The Multifaceted ATPase Inhibitory Factor 1 (IF1) in Energy Metabolism Reprogramming and Mitochondrial Dysfunction: A New Player in Age-Associated Disorders?Antioxid Redox Signal. 2022 Aug;37(4-6):370-393. doi: 10.1089/ars.2021.0137. Epub 2022 Jan 4. Antioxid Redox Signal. 2022. PMID: 34605675 Free PMC article. Review.
References
-
- Pullman ME, Monroy GC. A naturally occurring inhibitor of mitochondrial adenosine triphosphatase occurring adenosine inhibitor of mitochondrial triphosphatase. J Biol Chem (1963) 238:3762–9. - PubMed
-
- Aggeler R, Coons J, Taylor SW, Ghosh SS, García JJ, Capaldi RA, et al. . A functionally active human F1F0 ATPase can be purified by immunocapture from heart tissue and fibroblast cell lines. subunit structure and activity studies. J Biol Chem (2002) 277:33906–12. doi: 10.1074/jbc.M204538200 - DOI - PubMed
-
- García-Bermúdez J, Sánchez-Aragó M, Soldevilla B, del Arco A, Nuevo-Tapioles C, Cuezva JM. PKA phosphorylates the ATPase inhibitory factor 1 and inactivates its capacity to bind and inhibit the mitochondrial h+-ATP synthase. Cell Rep (2015) 12:2143–55. doi: 10.1016/j.celrep.2015.08.052 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

