The genome sequence of the segmented worm, Sthenelais limicola (Ehlers, 1864)
- PMID: 37469856
- PMCID: PMC10352628
- DOI: 10.12688/wellcomeopenres.18856.1
The genome sequence of the segmented worm, Sthenelais limicola (Ehlers, 1864)
Abstract
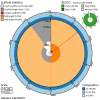
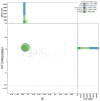
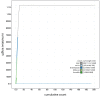
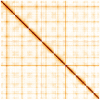
We present a genome assembly from an individual Sthenelais limicola (the segmented worm; Annelida; Polychaeta; Phyllodocida; Sigalionidae). The genome sequence is 1,131 megabases in span. Most of the assembly is scaffolded into nine chromosomal pseudomolecules. The mitochondrial genome has also been assembled and is 16.7 kilobases in length.
Keywords: Phyllodocida; Sthenelais limicola; chromosomal; genome sequence; segmented worm.
Copyright: © 2023 Darbyshire T et al.
Conflict of interest statement
No competing interests were disclosed.
Figures

References
-
- Barnich R, Fiege D: The Aphroditoidea (Annelida: Polychaeta) of the Mediterranean Sea. Abhandlungen der Senckenbergischen Naturforschenden Gesellschaft, 2003;559:1–167. Reference Source
-
- Chambers SJ, Muir AI: Polychaetes: British Chrysopetaloidea, Pisionoidea and Aphroditoidea. Synopses of the British Fauna (New Series). 1997;54:1–202. Reference Source
Grants and funding
LinkOut - more resources
Full Text Sources

