ATP-induced cross-linking of a biomolecular condensate
- PMID: 37480229
- PMCID: PMC11163290
- DOI: 10.1016/j.bpj.2023.07.013
ATP-induced cross-linking of a biomolecular condensate
Abstract
DEAD-box helicases are important regulators of biomolecular condensates. However, the mechanisms through which these enzymes affect the dynamics of biomolecular condensates have not been systematically explored. Here, we demonstrate the mechanism by which the mutation of a DEAD-box helicase's catalytic core alters ribonucleoprotein condensate dynamics in the presence of ATP. Through altering RNA length within the system, we are able to attribute the altered biomolecular dynamics and material properties to physical cross-linking of RNA facilitated by the mutant helicase. These results suggest that mutant condensates approach a gel transition when RNA length is increased to lengths comparable to eukaryotic mRNA. Lastly, we show that this cross-linking effect is tunable with ATP concentration, uncovering a system whose RNA mobility and material properties vary with enzyme activity. More generally, these findings point to a fundamental mechanism for modulating condensate dynamics and emergent material properties through nonequilibrium, molecular-scale interactions.
Copyright © 2023 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


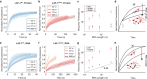

Update of
-
ATP-induced crosslinking of a biomolecular condensate.bioRxiv [Preprint]. 2023 Apr 18:2023.04.18.535486. doi: 10.1101/2023.04.18.535486. bioRxiv. 2023. Update in: Biophys J. 2024 Jun 4;123(11):1356-1366. doi: 10.1016/j.bpj.2023.07.013. PMID: 37131735 Free PMC article. Updated. Preprint.
Similar articles
-
Nonequilibrium phases of a biomolecular condensate facilitated by enzyme activity.bioRxiv [Preprint]. 2024 Aug 11:2024.08.11.607499. doi: 10.1101/2024.08.11.607499. bioRxiv. 2024. PMID: 39149291 Free PMC article. Preprint.
-
ATP-induced crosslinking of a biomolecular condensate.bioRxiv [Preprint]. 2023 Apr 18:2023.04.18.535486. doi: 10.1101/2023.04.18.535486. bioRxiv. 2023. Update in: Biophys J. 2024 Jun 4;123(11):1356-1366. doi: 10.1016/j.bpj.2023.07.013. PMID: 37131735 Free PMC article. Updated. Preprint.
-
Biomolecular condensates: It was RNA all along!Mol Cell. 2025 Feb 6;85(3):461-463. doi: 10.1016/j.molcel.2025.01.009. Mol Cell. 2025. PMID: 39919713
-
DEAD Box RNA Helicases: Biochemical Properties, Role in RNA Processing and Ribosome Biogenesis.Cell Biochem Biophys. 2024 Jun;82(2):427-434. doi: 10.1007/s12013-024-01240-w. Epub 2024 Mar 2. Cell Biochem Biophys. 2024. PMID: 38430409 Review.
-
Using quantitative reconstitution to investigate multicomponent condensates.RNA. 2022 Jan;28(1):27-35. doi: 10.1261/rna.079008.121. Epub 2021 Nov 12. RNA. 2022. PMID: 34772789 Free PMC article. Review.
Cited by
-
Structured protein domains enter the spotlight: modulators of biomolecular condensate form and function.Trends Biochem Sci. 2025 Mar;50(3):206-223. doi: 10.1016/j.tibs.2024.12.008. Epub 2025 Jan 17. Trends Biochem Sci. 2025. PMID: 39827079 Review.
-
Emerging biophysical principles of macromolecular phase separation.Biophys J. 2024 Jun 4;123(11):E1-E3. doi: 10.1016/j.bpj.2024.05.001. Epub 2024 May 17. Biophys J. 2024. PMID: 38761796 Free PMC article. No abstract available.
-
Quantifying surface tension and viscosity in biomolecular condensates by FRAP-ID.Biophys J. 2024 Oct 1;123(19):3366-3374. doi: 10.1016/j.bpj.2024.07.043. Epub 2024 Aug 8. Biophys J. 2024. PMID: 39113361
-
Nonequilibrium phases of a biomolecular condensate facilitated by enzyme activity.bioRxiv [Preprint]. 2024 Aug 11:2024.08.11.607499. doi: 10.1101/2024.08.11.607499. bioRxiv. 2024. PMID: 39149291 Free PMC article. Preprint.
References
-
- Brangwynne C., Tompa P., Pappu R. Polymer physics of intracellular phase transitions. Nat. Phys. 2015;11:899–904. doi: 10.1038/nphys3532. - DOI
-
- Guillén-Boixet J., Kopach A., et al. Franzmann T.M. RNA-Induced Conformational Switching and Clustering of G3BP Drive Stress Granule Assembly by Condensation. Cell. 2020;181:346–361.e17. https://www.sciencedirect.com/science/article/pii/S0092867420303421 - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

