Targeting MS4A4A on tumour-associated macrophages restores CD8+ T-cell-mediated antitumour immunity
- PMID: 37507218
- PMCID: PMC10715532
- DOI: 10.1136/gutjnl-2022-329147
Targeting MS4A4A on tumour-associated macrophages restores CD8+ T-cell-mediated antitumour immunity
Abstract
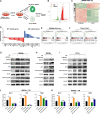
Objective: Checkpoint immunotherapy unleashes T-cell control of tumours but is suppressed by immunosuppressive myeloid cells. The transmembrane protein MS4A4A is selectively highly expressed in tumour-associated macrophages (TAMs). Here, we aimed to reveal the role of MS4A4A+ TAMs in regulating the immune escape of tumour cells and to develop novel therapeutic strategies targeting TAMs to enhance the efficacy of immune checkpoint inhibitor (ICI) in colorectal cancer.
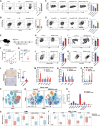
Design: The inhibitory effect of MS4A4A blockade alone or combined with ICI treatment on tumour growth was assessed using murine subcutaneous tumour or orthotopic transplanted models. The effect of MS4A4A blockade on the tumour immune microenvironment was assessed by flow cytometry and mass cytometry. RNA sequencing and western blot analysis were used to further explore the molecular mechanism by which MS4A4A promoted macrophages M2 polarisation.
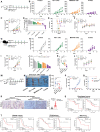
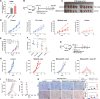
Results: MS4A4A is selectively expressed by TAMs in different types of tumours, and was associated with adverse clinical outcome in patients with cancer. In vivo inhibition of MS4A4A and anti-MS4A4A monoclonal antibody treatment both curb tumour growth and improve the effect of ICI therapy. MS4A4A blockade treatment reshaped the tumour immune microenvironment, resulting in reducing the infiltration of M2-TAMs and exhausted T cells, and increasing the infiltration of effector CD8+ T cells. Anti-MS4A4A plus anti-programmed cell death protein 1 (PD-1) therapy remained effective in large, treatment-resistant tumours and could induce complete regression when further combined with radiotherapy. Mechanistically, MS4A4A promoted M2 polarisation of macrophages by activating PI3K/AKT pathway and JAK/STAT6 pathway.
Conclusion: Targeting MS4A4A could enhance the ICI efficacy and represent a new anticancer immunotherapy.
Keywords: T lymphocytes; colorectal cancer; immunotherapy; macrophages.
© Author(s) (or their employer(s)) 2023. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: None declared.
Figures



Comment in
-
Targeting myeloid signalling pathways to unleash T cells.Gut. 2023 Nov 24;72(12):2223-2225. doi: 10.1136/gutjnl-2023-330706. Gut. 2023. PMID: 37863515 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous
