Proteomic Alteration in the Progression of Multiple Myeloma: A Comprehensive Review
- PMID: 37510072
- PMCID: PMC10378430
- DOI: 10.3390/diagnostics13142328
Proteomic Alteration in the Progression of Multiple Myeloma: A Comprehensive Review
Abstract
Multiple myeloma (MM) is an incurable hematologic malignancy. Most MM patients are diagnosed at a late stage because the early symptoms of the disease can be uncertain and nonspecific, often resembling other, more common conditions. Additionally, MM patients are commonly associated with rapid relapse and an inevitable refractory phase. MM is characterized by the abnormal proliferation of monoclonal plasma cells in the bone marrow. During the progression of MM, massive genomic alterations occur that target multiple signaling pathways and are accompanied by a multistep process involving differentiation, proliferation, and invasion. Moreover, the transformation of healthy plasma cell biology into genetically heterogeneous MM clones is driven by a variety of post-translational protein modifications (PTMs), which has complicated the discovery of effective treatments. PTMs have been identified as the most promising candidates for biomarker detection, and further research has been recommended to develop promising surrogate markers. Proteomics research has begun in MM, and a comprehensive literature review is available. However, proteomics applications in MM have yet to make significant progress. Exploration of proteomic alterations in MM is worthwhile to improve understanding of the pathophysiology of MM and to search for new treatment targets. Proteomics studies using mass spectrometry (MS) in conjunction with robust bioinformatics tools are an excellent way to learn more about protein changes and modifications during disease progression MM. This article addresses in depth the proteomic changes associated with MM disease transformation.
Keywords: mass spectrometry; multiple myeloma; post-translational modifications; proteomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
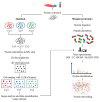



References
-
- Hari P., Mateos M.V., Abonour R., Knop S., Bensinger W., Ludwig H., Song K., Hajek R., Moreau P., Siegel D.S., et al. Efficacy and safety of carfilzomib regimens in multiple myeloma patients relapsing after autologous stem cell transplant: ASPIRE and ENDEAVOR outcomes. Leukemia. 2017;31:2630–2641. doi: 10.1038/leu.2017.122. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

