Pluripotency-Associated microRNAs in Early Vertebrate Embryos and Stem Cells
- PMID: 37510338
- PMCID: PMC10379376
- DOI: 10.3390/genes14071434
Pluripotency-Associated microRNAs in Early Vertebrate Embryos and Stem Cells
Abstract
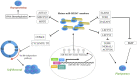
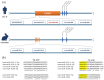
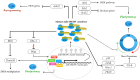
MicroRNAs (miRNAs), small non-coding RNA molecules, regulate a wide range of critical biological processes, such as proliferation, cell cycle progression, differentiation, survival, and apoptosis, in many cell types. The regulatory functions of miRNAs in embryogenesis and stem cell properties have been extensively investigated since the early years of miRNA discovery. In this review, we will compare and discuss the impact of stem-cell-specific miRNA clusters on the maintenance and regulation of early embryonic development, pluripotency, and self-renewal of embryonic stem cells, particularly in vertebrates.
Keywords: cell cycle regulators; embryonic development; embryonic stem cells; microRNA clusters.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

