Insights into the Evolution and Host Adaptation of the Monkeypox Virus from a Codon Usage Perspective: Focus on the Ongoing 2022 Outbreak
- PMID: 37511283
- PMCID: PMC10380431
- DOI: 10.3390/ijms241411524
Insights into the Evolution and Host Adaptation of the Monkeypox Virus from a Codon Usage Perspective: Focus on the Ongoing 2022 Outbreak
Abstract
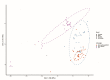
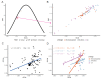
The exceptionally widespread outbreak of human monkeypox, an emerging zoonosis caused by the monkeypox virus (MPXV), with more than 69,000 confirmed cases in 100 non-endemic countries since 2022, is a major public health concern. Codon usage patterns reflect genetic variation and adaptation to new hosts and ecological niches. However, detailed analyses of codon usage bias in MPXV based on large-scale genomic data, especially for strains responsible for the 2022 outbreak, are lacking. In this study, we analyzed codon usage in MPXV and its relationship with host adaptation. We confirmed the ongoing outbreak of MPXVs belonging to the West Africa (WA) lineage by principal component analysis based on their codon usage patterns. The 2022 outbreak strains had a relatively low codon usage bias. Codon usage of MPXVs was shaped by mutation and natural selection; however, different from past strains, codon usage in the 2022 outbreak strains was predominantly determined by mutation pressure. Additionally, as revealed by the codon adaptation index (CAI), relative codon deoptimization index (RCDI), and similarity index (SiD) analyses, the codon usage patterns of MPXVs were also affected by their hosts. In particular, the 2022 outbreak strains showed slightly but significantly greater adaptation to many primates, including humans, and were subjected to stronger selection pressure induced by hosts. Our results suggest that MPXVs contributing to the 2022 outbreak have unique evolutionary features, emphasizing the importance of sustained monitoring of their transmission and evolution.
Keywords: codon usage bias; evolution; host adaptation; monkeypox virus; mutation pressure; natural selection.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Karagoz A., Tombuloglu H., Alsaeed M., Tombuloglu G., AlRubaish A.A., Mahmoud A., Smajlović S., Ćordić S., Rabaan A.A., Alsuhaimi E. Monkeypox (mpox) virus: Classification, origin, transmission, genome organization, antiviral drugs, and molecular diagnosis. J. Infect. Public Health. 2023;16:531–541. doi: 10.1016/j.jiph.2023.02.003. - DOI - PMC - PubMed
-
- Centers for Disease Control and Prevention 2022 Monkeypox Outbreak Global Map. [(accessed on 4 October 2022)]; Available online: https://www.cdc.gov/poxvirus/monkeypox/response/2022/world-map.html.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

