Sugar Starvation Disrupts Lipid Breakdown by Inducing Autophagy in Embryonic Axes of Lupin (Lupinus spp.) Germinating Seeds
- PMID: 37511532
- PMCID: PMC10380618
- DOI: 10.3390/ijms241411773
Sugar Starvation Disrupts Lipid Breakdown by Inducing Autophagy in Embryonic Axes of Lupin (Lupinus spp.) Germinating Seeds
Abstract
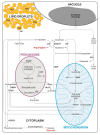
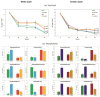
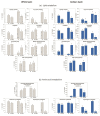
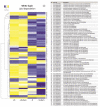
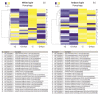
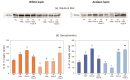
Under nutrient deficiency or starvation conditions, the mobilization of storage compounds during seed germination is enhanced to primarily supply respiratory substrates and hence increase the potential of cell survival. Nevertheless, we found that, under sugar starvation conditions in isolated embryonic axes of white lupin (Lupinus albus L.) and Andean lupin (Lupinus mutabilis Sweet) cultured in vitro for 96 h, the disruption of lipid breakdown occurs, as was reflected in the higher lipid content in the sugar-starved (-S) than in the sucrose-fed (+S) axes. We postulate that pexophagy (autophagic degradation of the peroxisome-a key organelle in lipid catabolism) is one of the reasons for the disruption in lipid breakdown under starvation conditions. Evidence of pexophagy can be: (i) the higher transcript level of genes encoding proteins of pexophagy machinery, and (ii) the lower content of the peroxisome marker Pex14p and its increase caused by an autophagy inhibitor (concanamycin A) in -S axes in comparison to the +S axes. Additionally, based on ultrastructure observation, we documented that, under sugar starvation conditions lipophagy (autophagic degradation of whole lipid droplets) may also occur but this type of selective autophagy seems to be restricted under starvation conditions. Our results also show that autophagy occurs at the very early stages of plant growth and development, including the cells of embryonic seed organs, and allows cell survival under starvation conditions.
Keywords: asparagine; embryo; iTRAQ; lipid droplet; lipophagy; peroxisome; pexophagy; proteomics; transcriptomics; ultrastructure.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Ratajczak W., Borek S., Podgorski A., Ratajczak L. Variability of globulin composition in cultivars and individually tested seeds of yellow lupin (Lupinus luteus L.) Acta Physiol. Plant. 1999;21:413–417. doi: 10.1007/s11738-999-0014-z. - DOI
-
- Borek S., Ratajczak W., Ratajczak L. Regulation of storage lipid metabolism in developing and germinating lupin (Lupinus spp.) seeds. Acta Physiol. Plant. 2015;37:119. doi: 10.1007/s11738-015-1871-2. - DOI
-
- Borek S., Galor A., Paluch E. Asparagine Enhances Starch Accumulation in Developing and Germinating Lupin Seeds. J. Plant Growth Regul. 2013;32:471–482. doi: 10.1007/s00344-012-9313-5. - DOI
-
- Gaufichon L., Reisdorf-Cren M., Rothstein S.J., Chardon F., Suzuki A. Biological functions of asparagine synthetase in plants. Plant Sci. 2010;179:141–153. doi: 10.1016/j.plantsci.2010.04.010. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

