A non-proteolytic release mechanism for HMCES-DNA-protein crosslinks
- PMID: 37519246
- PMCID: PMC10505908
- DOI: 10.15252/embj.2022113360
A non-proteolytic release mechanism for HMCES-DNA-protein crosslinks
Abstract
The conserved protein HMCES crosslinks to abasic (AP) sites in ssDNA to prevent strand scission and the formation of toxic dsDNA breaks during replication. Here, we report a non-proteolytic release mechanism for HMCES-DNA-protein crosslinks (DPCs), which is regulated by DNA context. In ssDNA and at ssDNA-dsDNA junctions, HMCES-DPCs are stable, which efficiently protects AP sites against spontaneous incisions or cleavage by APE1 endonuclease. In contrast, HMCES-DPCs are released in dsDNA, allowing APE1 to initiate downstream repair. Mechanistically, we show that release is governed by two components. First, a conserved glutamate residue, within HMCES' active site, catalyses reversal of the crosslink. Second, affinity to the underlying DNA structure determines whether HMCES re-crosslinks or dissociates. Our study reveals that the protective role of HMCES-DPCs involves their controlled release upon bypass by replication forks, which restricts DPC formation to a necessary minimum.
Keywords: DNA repair; DNA replication; DNA-protein crosslinks; HMCES; abasic (AP) sites.
© 2023 The Authors. Published under the terms of the CC BY NC ND 4.0 license.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

SRAP domain sequence alignment highlighting key active site residues in H. sapiens, X. laevis, S. cerevisiae and E. coli HMCES homologues (Cys2 = orange, Glu127 = red and His210 = green).
Proposed reaction mechanism of SRAP domain crosslinking to an AP site.
Crystal structure of HMCES' active site crosslinked to an AP site. PDB: 6OE7 (Halabelian et al, 2019). DNA is shown in grey. Active site residues are coloured as in (A). Interatomic distances (Å) are labelled.
Schematic of the assay shown in (E) and (G). HMCESFL and HMCESSRAP (WT or active site variants) were incubated for 1 h at 37°C with a Cy5‐labelled 30mer oligonucleotide containing either a dT or an AP site at position 15. Afterwards, non‐crosslinked HMCES was inactivated by heat denaturation at 60°C for 5 min. A second 6‐FAM‐labelled 30mer oligonucleotide containing an AP site was added and formation of 6‐FAM DPCs was assessed after an additional incubation for 120 min.
HMCESFL‐ and HMCESSRAP‐WT and HMCESSRAP‐C2S‐DPC formation with Cy5‐ and 6‐FAM‐oligonucleotides was analysed using denaturing SDS–PAGE. Incised DNA is caused by spontaneous hydrolysis of the AP site.
Quantification of DPC formation assays shown in (E), left panel: DPC formation to Cy5 oligonucleotide, right panel: DPC formation to 6‐FAM oligonucleotide.
DPC formation of HMCESSRAP‐WT and variants (E127A or H210A) with Cy5‐ and 6‐FAM‐oligonucleotides was analysed using denaturing SDS–PAGE. Incised DNA is caused by spontaneous hydrolysis of the AP site.
Quantification of DPC formation assays shown in (G), left panel: DPC formation to Cy5 oligonucleotide, right panel: DPC formation to 6‐FAM oligonucleotide.

Coomassie‐stained SDS–PAGE gel showing recombinant purified human HMCESFL‐WT, HMCESSRAP‐WT, HMCESSRAP‐C2S, HMCESSRAP‐R98E, HMCESSRAP‐E127A, HMCESSRAP‐H210A and HMCESSRAP‐R98E/E127A proteins used in this study.
DPC formation of HMCESFL and HMCESSRAP (WT or indicated variants). dU‐containing DNA (0.1 μM) was incubated alone or with UDG and increasing concentrations of HMCES (0.025, 0.05, 0.1, 0.2, 0.4, 0.8, 1.6 and 3.2 μM), as indicated for 1 h at 37°C prior to analysis by denaturing SDS–PAGE.
Quantification of HMCES‐DPC formation assays shown in (B).

Kinetics of DPC formation by HMCESSRAP (WT, R98E, E127A or H210A variants) to ssDNA, junction DNA and dsDNA. Corresponding reverse oligonucleotides were annealed to ssDNA to create DNA junction and dsDNA prior to adding HMCESSRAP. To ssDNA, a non‐complementary oligonucleotide was added as control. HMCESSRAP‐WT and variants were incubated with different DNA structures for the indicated amount of time at 37°C prior to separation by denaturing SDS–PAGE.
Quantification of DPC formation assays shown in (A)
Kinetics of DPC formation by HMCESSRAP‐WT to junction DNA and dsDNA containing a nick or a 4 nt‐gap. Corresponding reverse oligonucleotides were annealed to create junction DNA and dsDNA containing a nick or a 4 nt‐gap prior to adding HMCESSRAP‐WT. HMCESSRAP‐WT was incubated with the indicated DNA structures for the indicated amount of time at 37°C prior to separation by denaturing SDS–PAGE.
Quantification of DPC formation assays shown in (C).

DPC reversal kinetics of indicated variants in ssDNA, DNA junction and dsDNA. DPCs were pre‐formed in ssDNA before corresponding reverse oligonucleotides were annealed (for ssDNA reactions, a non‐complementary oligonucleotide was added). DPC reversal was then monitored after incubation for the indicated amount of time at 37°C using denaturing SDS–PAGE.
Quantification of DPC reversal assays using HMCESSRAP‐WT and variants shown in (A).

Fluorescence polarization measurements of Cy5‐labelled ssDNA (25 nM) incubated with increasing concentrations of HMCESSRAP‐WT, HMCESSRAP‐R98E, HMCESSRAP‐E127A or HMCESSRAP‐R98E/E127A for 20 min on ice prior to measuring fluorescence polarization.
Non‐covalent DNA binding of indicated HMCESSRAP variants was assessed by electrophoretic mobility shift assay. A Cy5‐labelled 30‐mer ssDNA (0.1 μM) was incubated with HMCESSRAP (0, 0.125, 0.5 or 2 μM) for 20 min at 4°C prior to analysis by native PAGE.
DPC reversal kinetics of indicated HMCESSRAP variants in dsDNA. A corresponding reverse oligonucleotide was annealed to HMCESSRAP‐DPCs, before incubation for the indicated amount of time at 37°C prior to separation by denaturing SDS–PAGE.
Quantification of DPC reversal assays shown in (C).

- A, B
Competition assay between HMCESFL and indicated HMCESSRAP variants. HMCESSRAP‐DPCs in ssDNA (A) or at ssDNA‐dsDNA junctions (B) were pre‐formed and then incubated with 10‐fold excess of HMCESFL for the indicated amount of time at 37°C prior to separation by denaturing SDS gel (upper panels).

- A, B
APE1 incision of an AP site protected by the indicated HMCESSRAP‐DPC variants at ssDNA‐dsDNA junctions (A) or within dsDNA (B). Free dU‐containing DNA was incubated alone or in the presence of UDG and HMCESSRAP for 1 h at 37°C. Next, corresponding reverse oligonucleotides were annealed to generate an ssDNA‐dsDNA junction (A) or dsDNA (B), and reactions were incubated alone or with APE1 for the indicated amount of time at 37°C prior to separation by denaturing SDS–PAGE.

- A
APE1 incision of an AP site in ssDNA, DNA junction and dsDNA. A Cy5‐labelled 30‐mer ssDNA was incubated alone or with UDG for 1 h at 37°C. Corresponding reverse oligonucleotides for DNA junction or dsDNA were annealed (for ssDNA, a non‐complementary oligonucleotide was added). Next, samples were incubated with APE1 for 18 h at 37°C before separation by denaturing SDS–PAGE.
- B
Coomassie‐stained SDS–PAGE gel showing recombinant purified bacterial YedK‐WT, YedK‐E105A, xl‐HMCES‐WT and xl‐HMCES‐E129A proteins used in this study.
- C, D
APE1 incision of an AP site protected by the indicated YedK‐WT‐DPC or YedK‐E105A‐DPC and xl‐HMCES‐WT‐DPC or xl‐HMCES‐E129A‐DPC at ssDNA‐dsDNA junctions (C) or within dsDNA (D). Free dU‐containing DNA was incubated alone or in the presence of UDG and YedK/xl‐HMCES for 1 h at 37°C. Next, corresponding reverse oligonucleotides were annealed to generate an ssDNA‐dsDNA junction (C) or dsDNA (D), and reactions were incubated alone or with APE1 for the indicated amount of time at 37°C prior to separation by denaturing SDS–PAGE.

- A
Schematic depiction of the NEIL3‐dependent repair of an AP‐ICL, a lesion that forms when an AP site reacts with a nucleobase of the opposing DNA strand forming a covalent crosslink (Price et al, 2014). In Xenopus egg extracts, such crosslinks are primarily unhooked by the NEIL3 glycosylase (Semlow et al, 2016), which yields an AP site leading to formation of an HMCES‐DPC.
- B–F
In the absence of SPRTN, the intact HMCES‐DPC is presumably bypassed by TLS and transferred into dsDNA. To test whether this triggers autorelease, we analysed the stability of DPCs formed by wild‐type and E129A‐mutated Xenopus laevis rHMCES‐3xFlag proteins during ICL repair in egg extract. pICL‐lacO AP was replicated in mock‐ or SPRTN‐depleted extracts (B) supplemented with WT or E129A rHMCES‐3xFlag. At the time points indicated, plasmid was recovered under stringent conditions, the DNA was digested and released proteins were separated by SDS–PAGE. HMCES‐DPCs were detected using an antibody raised against the SRAP domain that permits simultaneous monitoring of endogenous HMCES protein and the recombinant 3xFlag‐tagged HMCES (which migrates slower during SDS–PAGE due to the 3xFlag). In this experimental setup, the endogenous protein serves as a control for the effects of SPRTN depletion and autorelease. Like the endogenous HMCES (C), both WT (D) and E129A‐mutated rHMCES‐3xFlag (E) were stabilized by SPRTN depletion, implying that proteolysis is the dominant mechanism for removing HMCES‐DPC under these conditions. However, it was challenging to assess the relative behaviour of tagged WT and mutant protein because DPCs formed by the wild‐type recombinant protein (like those formed by the endogenous protein) are resolved slowly in SPRTN‐depleted extract (on the timescale of hours, somewhat slower than the timescale for observed for reversal in vitro). Additionally, the E129A‐mutated recombinant flag‐tagged protein crosslinked less efficiently than endogenous HMCES, making it difficult to detect even when present in large excess (F). We are, therefore, unable to determine from these data whether HMCES‐DPC reversal occurs during ICL repair in egg extract under the conditions tested. Orange dots denote non‐specific bands or bands corresponding to contaminating IgG.
- G
One explanation for the discrepancy in the degree of reversal observed between the in vitro reconstitution and egg extract systems could be that the in vitro reactions were all performed at 37°C, while replication in egg extracts must be performed at 20°C. Therefore, we assessed reversal of HMCESSRAP‐WT or ‐E127A‐DPCs in dsDNA at the indicated temperatures for the indicated amount of time before analysis by denaturing SDS–PAGE. Indeed, autocatalytic reversal was significantly delayed at 20°C.
- H
Quantification of DPC reversal assays using HMCESSRAP‐WT and ‐E127A shown in (G).
- I
The extracts used in the replication reactions shown in (J and K) were immunoblotted for SPRTN, Rev1 and HMCES.
- J
As an alternative additional strategy to determine whether reversal contributes to HMCES‐DPC resolution, we tested whether REV1 depletion results in stabilization of HMCES‐DPCs, reasoning that blocking TLS (and transfer of the DPC into dsDNA) may inhibit reversal. pICL‐lacO AP was replicated in mock‐, REV1‐, SPRTN‐ or REV1‐ and SPRTN‐depleted egg extracts, as indicated. At the indicated time points, plasmid was recovered under stringent conditions, the DNA was digested and released proteins were separated by SDS–PAGE. HMCES was detected by blotting. As expected, depletion of SPRTN alone resulted in a strong stabilization of HMCES‐DPCs. Depletion of REV1 alone did not stabilize HMCES‐DPCs, consistent with our data indicating that SPRTN represents the dominant mechanism for HMCES‐DPC resolution in egg extract. Surprisingly, when combined with SPRTN depletion, REV1 depletion partially suppressed the accumulation of HMCES‐DPCs. Superficially, this result is contrary to our expectations based on data presented in Fig 6. However, we interpret the result to indicate when the HMCES‐DPC is maintained at an ssDNA/dsDNA junction due to inefficient TLS, residual SPRTN or another protease can eventually degrade the HMCES‐DPC. Therefore, while these data do not provide evidence for HMCES‐DPC reversal during ICL repair in egg extract, they do reinforce the need for alternative removal mechanisms for HMCES‐DPCs present in dsDNA that are refractory to proteolysis.
- K
In parallel with the reactions shown in (J), pICL‐lacO AP was replicated in the indicated egg extracts supplemented with [α‐32P]dCTP. Replication intermediates were separated on a native agarose gel and visualized by autoradiography. SC, supercoiled. OC, open circular. Consistent with a TLS defect upon Rev1 depletion, we observed accumulation of gapped, circular plasmids in replication gels, implying that the HMCES‐DPC is maintained at an ssDNA‐dsDNA junction.

- A, B
Primer extension assay using Pol ζ‐Rev1. Fluorescently labelled primer‐template substrates containing an AP site at the indicated position were incubated alone or in the presence of HMCESSRAP‐WT or ‐E127A, recombinant human FANCJ and Pol ζ‐Rev1 as indicated for 2 h at 37°C prior to separation by denaturing UREA–PAGE. (A) Model of oligonucleotides. (B) Cy5 scan and 6‐FAM scan of denaturing UREA–PAGE.
- C
Quantification of Cy5 signals shown in (B).
- D
Quantification of 6‐FAM signals shown in (B).
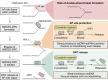

- A–C
HeLa T‐REx Flp‐In cells expressing the indicated doxycycline‐inducible HMCES variants with a C‐terminal mVenus‐3xFlag‐tag were grown in the presence of 1 μg/ml doxycycline, as indicated. Expression levels were analysed by Western blotting (A). Cell viability was determined using AlamarBlue cell viability assay (B), or crystal violet staining (C).
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

